Study design
This in-depth immunological analysis was conducted on samples obtained from KTRs selected from two clinical studies within the Dutch Renal Patients COVID-19 VACcination (RECOVAC) study. In the first study, performed between February 1st and May 31st, 2021, KTRs received two priming COVID-19 vaccinations. In the second study, which took place between October 20th, 2021, and February 5th, 2022, KTRs that did not seroconvert after the priming vaccine doses received additional COVID-19 vaccinations. For this study, KTRs that seroconverted after two priming COVID-19 vaccinations (primary responders) were selected and matched by age and sex to KTRs that seroconverted after a third or fourth vaccination (booster responders). Both studies were multicentre, conducted at four university medical hospitals in the Netherlands: Amsterdam UMC, UMC Groningen, Radboud UMC Nijmegen, and Erasmus MC Rotterdam. Approval for both studies was obtained from the Dutch Central Committee on Research Involving Human Subjects and the local ethics committees of the participating centres (Medische Ethische Toetsingscommissie AMC, Medische Ethische Toetsingscommissie Erasmus MC, Radboud University Medical Center Executive Board, Medisch Ethische Toetsingscommissie UMCG). This study was conducted in accordance with the Declaration of Helsinki, and written informed consent was received prior to participation.
Study participants and COVID-19 vaccination
All participants received two doses of the mRNA-1273 COVID-19 vaccine (Moderna Biotech Spain, S.L.) with a 28-day interval between doses, following the manufacturer’s guidelines. Blood samples were collected 28 days after the second vaccination to evaluate the immune response. For the booster study, individuals who did not seroconvert after receiving their second dose were included. These patients were randomly assigned in a 1:1:1 ratio to receive one of three booster vaccination regimens: a single dose of mRNA-1273 vaccine (100 μg, intramuscular), simultaneous administration of two doses of mRNA-1273 in both upper arms (2 × 100 μg, intramuscular), or a single dose of the Ad26.COV2-S vaccine (Janssen Biologics, Leiden, The Netherlands; 5 × 1010 viral particles, intramuscular). Similar to the priming study, blood samples were collected 28 days after the booster vaccination to assess immunogenicity. Throughout both studies, individuals with a history of COVID-19 (defined as confirmed by a positive SARS-CoV-2 test or the presence of nucleocapsid-specific antibodies) were excluded. Detailed inclusion and exclusion criteria can be found in the respective publications2,8, although importantly, all participants were clinically stable, at least one year post-transplantation, and free from rejection episodes in the preceding year. Data on the type of immunosuppressant therapy were available for every patient; detailed information on the therapy dose was missing for 12 of the primary responders. For this study, 40 age- and sex-matched KTRs from each study were included (Table 1). Immunoassays were conducted using the available samples, which led to differences in the number of patients included for each assay (Supplementary Fig. 1, Supplementary Table 1).
SARS-CoV-2 S1-specific IgG antibody response
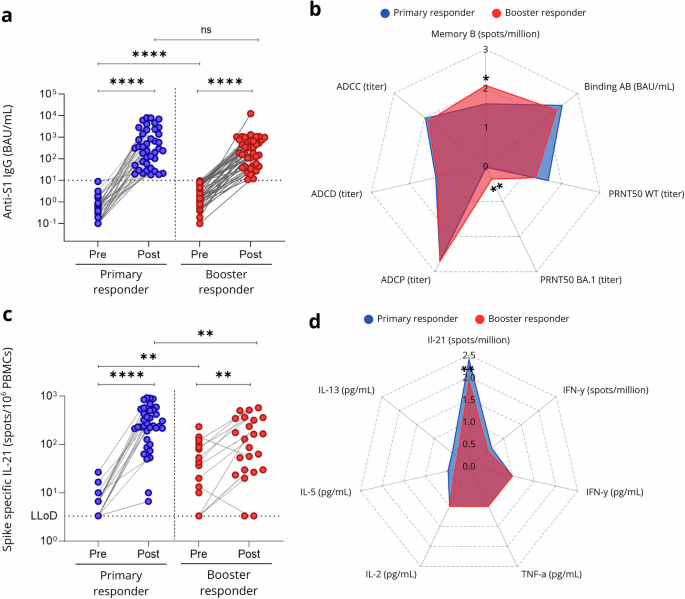
The SARS-CoV-2 S1-specific IgG antibody response in serum samples was measured at baseline and 28 days post priming or booster vaccination using a validated fluorescent bead-based multiplex immunoassay and expressed as BAU/mL. Patients were classified as seropositive or seronegative based on a threshold of 10 BAU/mL, determined by receiver operator curve analysis41,42.
Plaque reduction neutralization test (PRNT)
Neutralizing antibodies against SARS-CoV-2 were evaluated 28 days post-priming or booster vaccination using a plaque reduction neutralization test (PRNT), as previously described43,44. Heat-inactivated serum samples were tested against ancestral SARS-CoV-2 and Omicron (BA.1) variants.
Antibody-dependent cellular cytotoxicity
Antibody-dependent cell-mediated cytotoxicity (ADCC)-mediating antibodies were measured 28 days post-vaccination using an assay for NK cell degranulation as previously described43. Briefly, the NK92.05 cell line, genetically modified to express a high-affinity CD16 Fc receptor through a mutation at 176 V (NK92.05-CD16), was used. NK92.05-CD16 cells were cultured in Alpha-MEM supplemented with NaHCO3 (2.2 g/L, pH 7.2), 2-mercaptoethanol (0.0001 M), L-glutamine (200 mM), myo-inositol (0.2 mM), 10% horse serum, 10% fetal bovine serum, folic acid (0.004 mM), sodium pyruvate (1 mM), penicillin (100 IU/mL), streptomycin (100 μg/mL), and cultured cells were additionally supplemented with 100 IU/mL recombinant human IL-2 two times a week.
High-binding 96-well plates (Immunolon) were coated overnight at 4°C with 200 ng/well of trimeric prefusion biotinylated ancestral S protein (D614G, Sino Biological) or PBS (background control). Human heat-inactivated sera were 4-log serially diluted in PBS (1:40 to 1:163,840), and a positive control serum pool was included on each plate. After overnight serum incubation, plates were blocked, washed, and incubated with diluted serum at 37 °C for 2 hours. Next, plates were washed, and 100,000 NK92.05-CD16 cells were added with CD107aV450 (1:100, clone H4A3, BD), Golgistop (0.67 μL/mL, BD), and GolgiPlug (1 μL/mL, BD). Cells were incubated at 37°C for 5 hours, washed, and stained with CD56-PE (1:25, clone B159, BD) and LIVE/DEAD Fixable Aqua Dead Cell (AmCyan, Invitrogen, 1:100) at 4 °C for 30 minutes. Cells were fixed with Cytofix/Cytoperm (BD Biosciences) at 4 °C for 30 minutes. Activation of NK92.05-CD16 cells was measured in the CD56 + LIVE gate as the percentage of CD107a+ cells using a FACSLyric (BD). Percentages were adjusted by subtracting background levels measured on the PBS-coated well for the 1:10 dilution. ADCC induction was calculated from the standard curve and expressed as a 50% endpoint titer. When no ADCC induction was observed, the 50% titer was set at 40.
Antibody-dependent complement deposition
Antibody-dependent complement deposition (ADCD) was assessed 28 days post priming or booster vaccination on FluoSphere NeutrAvidin-labeled microspheres 1.0 µm beads (red, Invitrogen) coated with baculovirus-generated trimeric prefusion biotinylated ancestral (D614G) S protein (Sino Biological) as previously described45. For coating, FluoSphere beads were incubated with biotinylated protein in a 1:1 ratio at 37 °C for 2 hours. Following coating, S-coated FluoSpheres were washed 3 times with 5% BSA to block unbound regions on the bead. Beads were stored at 4 °C for up to 48 hours before use in the ADCD assay. Next, heat-inactivated human sera were added to S-coated FluoSphere beads in a 4-log dilution series starting at a 1:10 dilution and ranging to a 1:163,840 dilution, with the mixture then incubated at 37 °C for 2 hours. Next, a 1:50 dilution of guinea pig complement (Sanbio) in Gelatin Veronal Buffer supplemented with Ca2+ and Mg2+ (GVB++, CliniSciences BV) was added to each well and incubated for 15 minutes at 37 °C. To detect complement deposition, an anti-C3-FITC antibody (Fisher Scientific) at a 1:100 dilution was added and incubated for 20 minutes in the dark at 4 °C. C3 deposition on S-coated FluoSpheres was measured as FITC+ FluoSpheres by flow cytometry on a FACSLyric (BD). Background activation was subtracted from S-coated beads incubated with PBS. A serum pool from 19 healthy donors, as described above, was included in each experiment. ADCD titers relative to this standard were calculated and expressed as a 10% endpoint titer. When no ADCD was observed, the endpoint titer was set at 10.
Antibody-dependent cellular phagocytosis
ADCP was assessed 28 days post-priming or booster vaccination using the monocyte THP-1 cell line, as previously described45. THP-1 cells were cultured in RPMI 1640 with 0.05 mM 2-mercaptoethanol, 10% fetal bovine serum, 100 IU/mL penicillin, and 100 μg/mL streptomycin. For the phagocytosis assay, 1.0 μm FluoSphere NeutrAvidin-labeled beads (Invitrogen) were coated with baculovirus-produced biotinylated ancestral (D614G) prefusion trimeric S protein (Sino Biological) by incubating 5 μL beads with 5 μg protein at 37°C for 2 hours. Beads were washed three times with 5% BSA/PBS, resuspended in 0.01% BSA/PBS, and 1.8 × 10^8 coated beads were added per well in a 96-well plate. Serum was serially diluted 1:40 to 1:163,840 in PBS and incubated with beads at 37°C for 2 hours. Then, 50,000 THP-1 cells were added and incubated overnight at 37°C. Cells were stained with CD32-APC (BD) and LIVE/DEAD Fixable Aqua (Invitrogen) and analyzed by flow cytometry (FACSLyric, BD). Phagocytosis was quantified as gMFI of PE-positive cells within the CD32 + LIVE gate (20,000 cells/sample). Background was determined using beads incubated with PBS instead of serum. A standard curve from pooled serum of 19 healthy donors was used to calculate 50% endpoint ADCP titers; undetectable samples were assigned a titer of 40.
SARS-CoV-2-specific memory B cell response
The SARS-CoV-2-specific memory B cell response was assessed at baseline and 28 days post priming or booster vaccination, using a commercial B cell ELISpot (U-CyTech biosciences) according to the manufacturer’s instructions46. Peripheral blood mononuclear cells (PBMCs) were isolated from heparinized blood via Ficoll Paque and stored at −80°C until analysis. Thawed PBMCs (2,000,000 per well) were polyclonally activated with IL-2 and R848 for 7 days. Subsequently, 100,000 cells for SARS-CoV-2-specific and 10,000 for total IgG-producing cells were transferred to a 96-well plate pre-coated with a capture antibody for IgG and incubated for 20 hours. Detection was performed using biotinylated antibodies for IgG and SARS-CoV-2 S, followed by horseradish peroxidase and 3-Amino-9-ethylcarbazole staining. Spots were counted using an ELISpot reader (Bioreader 6000-V; Bio-Sys). The memory B cell response was quantified as the number of SARS-CoV-2-specific IgG-producing cells (spots) per 106 PBMCs. The lower limit of detection (LLoD) of this response was 10 B cell spots per 106 PBMCs.
SARS-CoV-2-specific IFN-γ and IL-21 memory T cell responses
SARS-CoV-2-specific IFN-γ and IL-21 memory T cell responses were evaluated similarly at baseline and 28 days post priming or booster vaccination using commercial IFNγ (Mabtech) and IL-21 (U-CyTech biosciences) ELISpot assays, according to the manufacturer’s guidelines20,46. Thawed PBMCs (200,000 (IFNγ) or 300,000 (IL-21) per well) were added to a 96-well plate pre-coated with an IFNγ or IL-21 capture antibody. Cells were stimulated with SARS-CoV-2 antigens for 20 (IFNγ) or 44 (IL-21) hours using S1 and S2 overlapping peptide pools (15-mer overlapping by 11, of 0.5 μg/mL for each peptide, JPT Peptide Technologies, PM-WCPV-S-1 & PM-WCPV-S-2). Detection involved biotinylated IFNγ or IL-21 antibodies, followed by streptavidin-horseradish peroxidase incubation. To visualize the spots, filtered TMB substrate was used for the IFNγ ELISpot and 3-Amino-9-ethylcarbazole for the IL-21 ELISpot. Spots were counted with an ELISpot reader (IFNγ: AID ELISpot reader; IL-21: Bioreader 6000-V; Bio-Sys). The IFNγ/IL-21 memory T cell response was expressed as the number of SARS-CoV-2-specific cytokine–producing cells (spots) per 106 PBMCs. For each stimulation, the average of the DMSO negative control was subtracted. The LLoD for was 3.3 IL-21 spots per 10⁶ PBMCs, and 5 IFN-γ spots per 10⁶ PBMCs at baseline and/or 28 days post-vaccination.
SARS-CoV-2-specific T cell cytokine responses
The human Th cytokine panel kit (LEGENDplex, BioLegend, CA, USA) was utilized to quantify cytokines in JPT antigen-stimulated PBMC culture supernatants from the samples of the IL-21 ELISpot 28 days post-vaccination. This panel included interleukin (IL)-2, IL-4, IL-5, IL-6, IL-9, IL-10, IL-13, IL-17A, IL-17F, IL-22, IFN-γ, and TNF-α19,47. Culture supernatants were thawed on ice, centrifuged, and prepared as twofold dilutions. These samples were then incubated with monoclonal capture antibody-coated beads for 2 hours. After washing, the beads were incubated with biotin-labeled detection antibodies for one hour, followed by streptavidin-PE for 30 minutes. Following staining, beads were acquired on a BD FACSCanto™ II with BD FACSDiva™ software (BD). Data acquisition and analysis were conducted using LEGENDplex V8.0 software (BioLegend). Cytokine concentrations were determined by assessing the streptavidin-PE signal against a freshly prepared standard curve, expressed as picograms per milliliter (pg/mL) after subtracting the background signal from the negative control. If subtraction resulted in a negative value, it was adjusted to 0 pg/mL.
T cell spectral flow cytometry
To obtain an in-depth insight into the phenotype of (SARS-CoV-2-specific) T cells, a spectral flow cytometry panel was employed on PBMC stimulated with overlapping S peptide pools. PBMCs were thawed and resuspended at 4 × 106 cells/mL. In each experimental batch, a healthy reference PBMC sample from the same individual was included to control for batch effects. After one hour of resting, 1 × 106 cells were seeded in polypropylene tubes for stimulation. To each tube, 2 µL of anti-CD40 antibody (Miltenyi Biotec, 130-094-133) was added to limit B cell-mediated CD154 (CD40L) internalization upon binding to CD40. Cells were stimulated with SARS-CoV-2 S peptide pools PM-WCPV-S-1 & PM-WCPV-S-2 (JPT Peptide Technologies) at 0.5 μg/mL or equimolar amounts of DMSO (negative control). Cells were cultured for 20 hours at 37 °C, 5% CO2. After stimulation, cells were prepared for staining. The panel was based on a panel used previously in the RECOVAC cohort48, optimized for staining index in this experimental setting (Supplementary Table 2). PBMCs were washed with PBS before staining for 30 minutes with Zombie UV dye (Biolegend, CA, USA) to detect dead cells. After washing with 3% FCS in PBS, 10 µL of Brilliant Stain Buffer Plus (BD Biosciences) and 5 µL of True stain monocyte blocker were added to each tube. Antibodies were consecutively added to the cells, without washing steps. First, γδTCR, CXCR3, and CD197 (CCR7) antibodies were added one by one, with ten minutes of incubation between each addition. Afterwards, all antibodies labelled with a BUV, BB, or BV fluorochrome were added individually, but without additional incubation time between each antibody. Finally, the remaining antibodies were added simultaneously. After 30 minutes incubation, cells were washed, fixed in 1% PFA, and measured on the Aurora spectral flow cytometer (Cytek Biosciences).
The phenotype of CD4, CD8, and γδTCR T-cells, as well as SARS-CoV-2-specific T-cells, was analysed by conventional gating (Kaluza V2.1 (Beckman Coulter)) and through Uniform Manifold Approximation and Projection (UMAP) analysis (FCS Express 7 software (De Novo Software)). Conventional gating was applied to identify and quantify commonly known T cell phenotypes and activation-induced marker (AIM)+ cells. AIM + CD4 T cells were defined as CD134 (OX40) and CD154 (CD40L) positive, AIM + CD8 T cells as CD69 and CD137 positive. CD45RA expression was monitored with special care to check for individuals with a C77G polymorphism, preventing normal interpretation of CD45RA expression49. In the UMAP analyses, for each unstimulated and stimulated sample, single, LIVE CD3+T cells were identified; these data were merged into a unified file with a unique identifier for each sample. An interval down-sampling method was used to select 10,000 cells for each sample; this cell number was achievable in more than 95% of the samples, with the remaining samples having slightly fewer cells included. The UMAP for total T cells was generated using 18 markers, excluding CD3, markers of the AIM assay (CD134, CD154, CD69, and CD137), and markers with a low (< 1% positive) expression (CD366, CD152, CD223, and CD279). The UMAP was constructed following a scaling procedure to standardize the expression values, with the settings: number of neighbors = 50, minimal distance = 0.1, and number of iterations = 2000. To check whether batch differences were still present after correcting the unmixing between batches based on the bridging healthy samples, UMAPs were compared for each batch, and no obvious differences were identified. Finally, two additional UMAPs were designed specifically for pre-gated CD4 T cells and CD8 T cells. A new unified file was made in which 7500 cells or 5000 cells were selected for each sample (CD4 and CD8 T cells, respectively), which was also reached for 95% of the samples. The same settings as for the total T cell UMAPs were applied; additionally, CD4, CD8, and γδTCR were excluded as generating markers.
Statistical analysis
First, we described the baseline characteristics of the primary and booster responder groups. Categorical variables were reported as numbers (percentages), and differences between groups were assessed using Fisher’s exact test. Continuous variables were presented as medians (interquartile ranges), with differences evaluated using the Kruskal-Wallis test for the alternative vaccination strategies. Second, the levels of S1-specific binding IgG, neutralizing, ADCC, ADCP, and ADCD antibodies were reported. Group differences were assessed using the Mann-Whitney U test or Pearson Chi-square test, depending on the data type and distribution. Additionally, the Wilcoxon Signed Rank test was used to investigate within-group differences. To visualize overall patterns of immune response quality, group mean values of log₁₀(x + 1)-transformed B cell and T cell measurements were displayed in spider plots. Third, Spearman’s correlation coefficient was calculated to explore relationships between S1-specific binding antibodies, neutralizing, ADCC, ADCP, ADCD antibodies, and T cell responses, including IL-21, IL-5, IL-13, IL-2, IFN-γ, TNF-α, and AIM + CD4 and CD8 cells. Correlation matrices were generated to examine these associations while handling pairwise complete observations. This approach ensured robust handling of missing data while maintaining statistical rigor. Fourth, principal component analysis (PCA) was performed to identify patterns in the antibody and T cell responses and reduce dimensionality. Prior to PCA, missing data were handled using multiple imputation via the ‘mice’ package with predictive mean matching (PMM). Five imputed datasets were generated, and one completed dataset was selected for further analysis. A log transformation (log10(x + 1)) was applied to the imputed data to normalize distributions. PCA was then conducted on the transformed dataset with unit variance scaling (standardization) to ensure equal contributions of all variables. The PCA results, including loadings and scores, were visualized using scree plots, score plots, biplots, and variable contribution plots. Additionally, individuals were visualized with confidence ellipses grouped by responder categories to explore clustering patterns in both antibody and T cell responses. Finally, for the spectral flow cytometry analyses, the two responder groups were compared by Mann-Whitney U tests and stimulated and unstimulated samples with the Wilcoxon signed rank test. Based on the CD4 UMAP, three main clusters of AIM + CD4 T cells were identified, which were quantified for each unstimulated and stimulated sample as a percentage of CD4 T cells. Associations of these clusters with the ELISpots and antibody concentrations were assessed by Spearman’s correlation. For the comparison of the total T cell compartment, our comparison was focused on subsets that were driving the UMAP clustering. These subsets were subsequently manually gated and quantified in unstimulated samples. Statistical analyses were performed using GraphPad Prism software version 9.1.2 and RStudio software version 4.0.5. A p-value < 0.05 was considered statistically significant.