Reagents
Erastin (a ferroptosis activator), Ferrostatin-1(Fer-1, a ferroptosis inhibitor), Anti-VEGFA (Ranibizumab), iFSP1 (a selective and glutathione-independent inhibitor of ferroptosis suppressor protein 1), and ISRIB (a reagent that can effectively reverse the phosphorylation of eukaryotic initiation factor 2α) were obtained from Selleck Chemicals (Houston, TX, USA). Vitamin K1, warfarin, and warfarin sodium were purchased from Aladdin Scientific (Shanghai, China). DMEM medium, fetal bovine serum (FBS), penicillin/streptomycin, bovine serum albumin (PBS), Hanks’ Balanced Salt Solution (HBSS), and Live/dead Kit were purchased from Invitrogen (Carlsbad, CA, USA). Paraformaldehyde (PFA), bovine serum albumin (BSA), and Triton X-100 were obtained from Beyotime (Jiangsu, China). The Iron Assay Kit, GSH/GSSG Quantification Kit, Cell Counting Kit-8 (CCK-8) reagent, Malondialdehyde (MDA) Assay Kit, Lipid Peroxidation Probe BDP 581/591 C11, Highly Sensitive DCFH-DA-ROS Assay Kit, and JC-1 MitoMP Detection Kit were purchased from Dojindo (Kumamoto, Japan).
Ethics statement
All the rearing and experimental procedures adhered to the Association for Research in Vision and Ophthalmology Statement and the ARRIVE guidelines and were approved by the Ethical Committee on Animal Experiments of Animal Care Committee of Zhongshan Hospital, Fudan University (No. 2019-285), Shanghai, China.
Animals
Male C57BL/6J mice (6–8 weeks old; 20 ~ 25 g) were purchased from SLAC Laboratory Animal (Shanghai, China). The animals underwent a 1-week acclimation period prior to any experimental procedures. All animals were housed in standard mouse cages (370 × 270 × 170 mm) with 3 mice per cage, provided with standard food and clean water ad libitum. All mice were monitored twice daily for survival throughout the experiment.
Laser-induced CNV model establishment and treatment
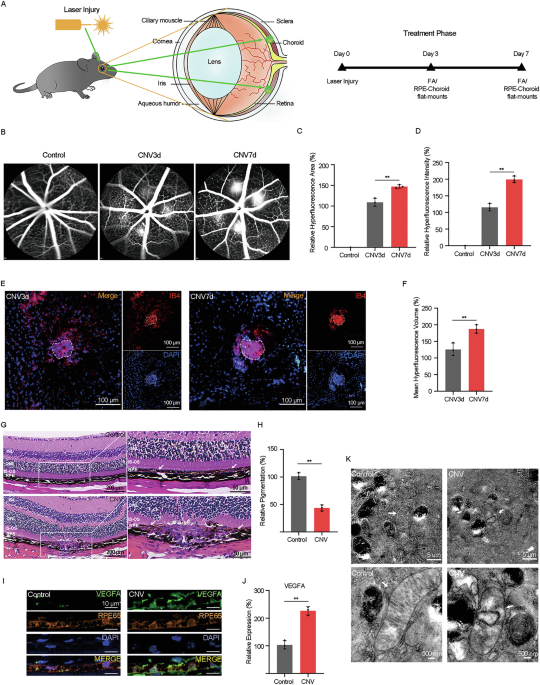
CNV was induced by photocoagulation, following established protocols [38]. Briefly, mice were anesthetized with 2% sodium pentobarbital (30 mg/kg; Sigma-Aldrich, St. Louis, MO, USA), and anesthetic depth was evaluated by the loss of the righting reflex. Pupil dilation was achieved using 1% tropicamide (Alcon Laboratories, Ft. Worth, TX, USA). Laser photocoagulation (532 nm, 120 mW, 50-μm spot size, 100 ms) was performed bilaterally using a diode-pumped solid-state laser (VISULAS 532 s; Carl Zeiss Meditec, Jena, Germany). 20 laser spots per eye were applied around the optic nerve for the western blot, while 4-6 laser spots per eye were applied for other experiments. Following laser irradiation, mice were placed on a heating plate at 37 °C until they regained consciousness. After a predetermined time, mice were euthanized for subsequent experiments by intraperitoneal injection of an overdose of 2% sodium pentobarbital (200 mg/kg) following spectral domain optical coherence tomography and fundus fluorescein angiography. The eyes were enucleated, and the neuroretina underlying the RPE were carefully separated to form eye cups.
Mouse RPE cells were separated from other structures through enzymic digestion (hyaluronidase and dispase; Medchem Express, Monmouth Junction, NJ, USA) as previous described [6]. Specifically, eye cups were digested with enzymic at 37 °C for 30 min, then washed with BSA/PBS, followed by mechanical dissection. The RPE cells identification was performed by Western blot (Fig. 2A).
In vivo research design and drug management
In the first in vivo experiment, C57BL/6J mice were randomly assigned to three groups: (1) Control group (n = 6), (2) CNV3d group (n = 6), and (3) CNV7d group (n = 6), based on the day of sacrifice after laser irradiation.
In the second in vivo experiment, C57BL/6J mice were divided into three groups: (1) mice with intravitreal injection of 1 μl of DMSO (Control group, n = 6); (2) mice with intravitreal injection of 1 μl of DMSO one day after laser irradiation (CNV group, n = 6); (3) mice with intravitreal injection of 1 μl of Fer-1(30 μM) one day after laser irradiation (Fer-1 group, n = 6). After one week, mice were used for subsequent experiments.
In the third in vivo experiment, C57BL/6 J mice were divided into five groups: (1) mice with daily intraperitoneal injection of 200 μl corn oil (Control group, n = 6); (2) mice with daily intraperitoneal injection of 200 μl corn oil after laser irradiation (CNV group, n = 6); (3) mice with intraperitoneal injection of Vitamin K1 (25 mg/kg) after laser irradiation (VK group, n = 6); (4) mice with warfarin sodium through bottled drinking water (0.33 mg/ml water) after laser irradiation (VKA group, n = 6); (5) mice with intraperitoneal injection of Vitamin K (20 mg/kg) and warfarin sodium through bottled drinking water (0.33 mg/ml water) after laser irradiation (VK + VKA group, n = 6). After one week, mice were used for subsequent experiments.
Spectral domain optical coherence tomography (SD-OCT) and fundus fluorescein angiography (FFA)
SD-OCT and FFA assessments were conducted at predetermined time points before sacrifice. Prior to examination, mice were deeply anaesthetized, and their pupils were dilated using tropicamide phenylephrine ophthalmic solution (Mydrin; Santen Pharmaceutical, Osaka, Japan). Positioned on the platform, the ocular fundus of the mice was sequentially examined using the SD-OCT system (Heidelberg Engineering, Heidelberg Germany). Subsequently, 50 μl of 10% fluorescein sodium solution (0.1 ml/kg; Alcon Laboratories, Ft. Worth, TX, USA) was administered via intraperitoneal injection. The ocular fundus was then consecutively imaged using a commercial digital fundus camera (Heidelberg Retina Angiograph, Heidelberg, Germany). The fluorescein leakage level was measured by analyzing fluorescence areas and intensity using ImageJ software.
RPE-choroid flat-mounts
At predetermined time points, mice were euthanized by intraperitoneal injection of an overdose of 2% sodium pentobarbital (200 mg/kg). Eyeballs were extracted for fluorescent-labeled isolectin staining of the CNV lesions. All eyeballs were carefully enucleated, fixed with 4% PFA, and then incubated with a blocking solution comprising 5% Goat serum and 0.3% Triton X-100 in PBS. Then, the eyecups were stained with FITC-conjugated Bandeiraea simplicifolia isolectin B4 (IB4; 1:100; VectorLabs, Newark, CA, USA) in blocking buffer at 4 °C for 8 h, and the nuclei were stained with DAPI (Invitrogen) for 15 min, shielded from light. Following staining, the RPE/choroid/sclera complex was carefully dissected into four incisions and mounted flat on a glass slide with the RPE side facing upward. All flat-mounts images were captured usings a confocal laser scanning microscope (FV3000; Olympus, Tokyo, Japan), and the area of CNV-related fluorescence was quantified using ImageJ software.
Hematoxylin – eosin (HE) and immunofluorescence (IF) staining
At the predetermined time points, the mice were euthanized, and their eyeballs were carefully extracted and immediately stored in FAS eyeball fixative solution (Servicebio, Wuhan, China). After dehydration in a series of ethanol solutions, the eyeballs were embedded in paraffin. Subsequently, tissue blocks embedded in paraffin were sectioned to a thickness of approximately 5 μm. HE staining was performed on the sections using hematoxylin-eosin. As described previously [6], IF slices were washed with PBS and fixed with goat serum, and then incubated with the antibodies listed in Supplementary Table S1 for 8 h at 4 °C. Images were obtained from at least three regions of each slice using a Nikon optical microscope (Tokyo, Japan). The primary antibody was detected using a fluorescent secondary antibody (Alexa Fluor 488/594; 1:500; Invitrogen) for 1 h at 37 °C. DAPI (Invitrogen) were used to stained the nuclei.
Transmission electron microscopy (TEM)
Within 2 min of eyeball extraction, a small piece of retina was removed from the posterior pole and fixed at 4 °C with 2.5% glutaraldehyde phosphate (0.1 M, pH 7.4) (Science services, Munich, Germany) for 2 h, followed by fixation with 2% osmium tetroxide for another 2 h. Then samples were dehydrated and embedded in Epon812 (Merck, Darmstadt, Germany). Subsequently, tissues were sectioned to a thickness of 60 nm, stained with lead citrate and uranyl acetate, and examined using transmission electron microscopy. Pictures were obtained using TEM microscopy (FEI, Hillsboro, OR, USA).
Iron ion detection
RPE cells were collected following the manufacturer’s instructions of the Iron Assay Kit. The absorbance at a wavelength of 593 nm was measured using a microplate reader (Multiskan skyhigh, Thermo Fisher Scientific, Waltham, MA, USA). The concentration of ferrous iron was calculated based on a standard curve.
GSH/GSSG and MDA measurement
After collecting the RPE cells, they were lysed by ultrasonication at 0 °C for 20 min, followed by centrifugation at 4 °C at 15,000 × g for 10 min. The cleared supernatant was used for subsequent detection. The GSH content and GSH/oxidized glutathione (GSSG) ratio were measured using a GSH/GSSG Quantification Kit according to the manufacturer’s instructions. The Malondialdehyde (MDA) content was assessed using an MDA Assay Kit following the manufacturer’s instructions. Total protein concentration was determined using a BCA Protein Assay Kit (Thermo Fisher Scientific).
Western blot analysis
Total protein was extracted using RIPA buffer (Invitrogen) containing 1% proteinase inhibitors (New Cell & Molecular Biotech, Suzhou, China) on ice, and the lysates were sonicated. Protein concentrations were determined using a BCA Protein Assay Kit.
The lysate proteins were isolated by SDS-PAGE gels (Bio-Rad, Hercules, CA, USA) and transferred onto PVDF membranes (Merck). After blocking with 5% BSA for 1 h, the membranes were incubated at 4 °C with different primary antibodies listed in Supplementary Table S2. Subsequently, the membranes were washed three times with TBST (pH 7.4, 10 mM Tris–HCl, 150 mM NaCl, and 0.1% Tween 20) and then incubated with horseradish peroxidase conjugated secondary anti-rabbit/mouse antibodies (1:5,000; Cell Signaling Technology, Danvers, MA, USA). Protein signals was viewed using an ECL Plus Western Blot Detection Kit (Tanon, Shanghai, China).
Cell culture and drug treatment
The human RPE cell line, ARPE-19 cells, obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China), and primary HUVEC provided by ScienCell Research Laboratories (Carlsbad, CA, USA) at passages 3–8 were used. The culture medium comprised DMEM, 10% FBS, and 1% penicillin/streptomycin, with medium changes every 2 days prior to conducting experiments.
Erastin (dissolved in DMSO, 1, 3, 5, 10, 20 μM) was used to induce ferroptosis cell death. Fer-1 (dissolved in DMSO, 10 μM) and Vitamin K1 (dissolved in ethyl alcohol, 0.1, 1, 3, 5, 10, 20, 25, 30, 50, 100 μM) were added to the culture medium 1 h before treating with ferroptosis activator. Warfarin (dissolved in DMSO, 1, 5, 10 μM), iFSP1 (dissolved in DMSO, 10 μM), and ISRIB (dissolved in DMSO, 10 μM) were added during cell seeding.
In the HUVEC conditioned culture experiments, the conditioned medium consisted of ARPE-19 cell supernatant. Specifically, ARPE-19 cells were cultured on the upper chambers of 0.4 μm-pore 6-well Costar Transwells for over 42 days to confluence and mature [37], following stimulation experiments as indicated for another 24 h. Then the culture medium was replaced with fresh medium and the supernatant was collected from the basolateral Transwell compartments after 24 h. Then, the scratch migration and tube formation assays of HUVEC were performed following incubation with the conditioned medium for 24 h. The concentration of FBS in the ARPE-19 culture medium for the scratch migration assay was 2%.
CCK-8 assays
The CCK-8 kit was used to evaluate the viability of ARPE-19 cells. Cells (2 × 104 cells/well) were seeded in 48-well plates and treated as indicated. Subsequently, the cells were digested using pancreatic enzymes, centrifuged, and suspended in equal volumes, and seeded in equal volumes into 96-well plates. Then, 10 µL CCK-8 reagent was added to each well and incubated at 37 °C for 4 h. The absorbance at 450 nm was measured using a microplate reader (ELX800, BioTek, Suwanee, GA, USA). Five independent experiments were performed.
Live/dead staining
A Live/dead Kit was used to stain the ARPE-19 cells (4 × 104 cells per well) cultured in a 24-well plate for the determination of viability, as directed by manufacturer’s instructions. At the determined time, the cells were incubated with Calcein AM and PI reagents for 15 min, and pictures were captured using a fluorescence microscope (Nikon).
ROS assays
The ROS level was evaluated using a highly Sensitive DCFH-DA-ROS assay kit following the manufacturer’s instructions. In brief, cells were washed twice with HBSS and then incubated with the highly sensitive DCFH-DA dye working solution for 30 min. Fluorescence signals were obtained using confocal laser scanning microscopy (FV3000, Olympus).
Lipid peroxidation assessment
ARPE-19 cells were treated as indicated and incubated for 1 h, protected from light, with a 50 μM Lipid Peroxidation Probe BDP 581/591 C11. After washing twice with PBS to remove excess probes, a small amount of serum-free medium was added to cover the cells in the dish. Fluorescence signals were obtained using confocal laser scanning microscopy (FV3000, Olympus).
Mitochondrial membrane potential detection
Mitochondrial membrane potential was evaluated using a JC-1 MitoMP Detection Kit according to the manufacturer’s instructions. After 4 h of erastin treatment, JC-1 solution was added to the culture medium to a final concentration of 4 μM and then incubated at 37 °C for 30 min. Following washing twice with HBSS, the cells were incubated in Imaging Buffer Solution, and fluorescence signals were obtained using confocal laser scanning microscopy (FV3000, Olympus).
RNA extraction and quantitative polymerase chain reaction (qPCR)
RNA extraction and quantitative PCR were performed as previously described [39]. Total RNA from ARPE-19 cells (5 × 105) in 6-well plates was extracted using TRIzol reagent (Takara, Kyoto, Japan). Reverse transcription of RNA into cDNA was carried out using PrimeScript RT master mix (Takara). To determine mRNA expression levels, real-time qPCR was performed using a Real-Time PCR Detection System (Applied Biosystems, Foster, CA, USA) according to the manufacturer’s instructions. The relative gene expression was analyzed by the 2 − ΔΔCt method. The relative mRNA expression is expressed as the fold change relative to GAPDH expression to ensure accurate gene quantification. Primer sequences were listed in Supplementary Table S3.
ELISA quantification
ARPE-19 cells were cultured on the upper chambers of 0.4 μm-pore 6-well Costar Transwells for over 42 days to confluence and mature, and then treated as indicated for 24 h. The culture medium was refreshed 24 h before the supernatant was collected from the basolateral Transwell compartments. The concentration of VEGFA was measured by an ELISA quantification kit (ABclonal, Wuhan, China).
Scratch migration assay
When HUVEC cells reached 100% confluence in 6-well plates, a wound injury was created by scratching the cell monolayer using disposable 200 μl pipette tips. Then, PBS was used to wash the cell debris away, and the cells were incubated in the corresponding ARPE-19 conditioned medium. A Nikon digital camera screened all the pictures of wound healing assays. Images of wound healing were taken per 12 h, and we chose the images of 0 h, 12 h, 24 h to assess the lateral migration ability of cells. Experiments were repeated three times, and the final scratch fusion area was calculated using ImageJ software.
Tube formation assay
The tube formation assay was conducted following established protocols [40]. Matrigel (Corning, USA) was pre-coated onto 96-well plates, and resuspended HUVEC cells were seeded into each well with conditioned medium composed of ARPE-19 cell supernatant treated as indicated. After incubated for 8 h, five random fields from each well were imaged using a microscope (Nikon). The experiments were repeated three times, and tube formation ability was quantified by measuring the number of nodes, tubes, and total segment length using ImageJ software.
RNA-sequencing and analysis
Total RNA of ARPE-19 cells was extracted and reverse-transcribed into cDNA using TRIzol reagent and PrimeScript RT master mix, as previously described [6]. The ligated products were amplified with PCR following a series of pretreatments, with an average insert size of 300 ± 50 bp for the final cDNA library. The 2 × 150 bp paired-end sequencing (PE150) was performed on an Illumina Novaseq™ 6000 (LC-Bio Technology Co., Ltd., Hangzhou, China). The RNA sequencing data has been deposited in the National Center for Biotechnology Information’s Gene Expression Omnibus (GEO) database.
siRNA transfection
ARPE-19 cells were seeded in 6-well plates for 24 h and transfected with siRNA (Gene Pharma, Shanghai, China) following the manufacturer’s instructions. The siRNA sequences used in this study were siATF4 (Qiagen, SI03019345), siVKORC1L1(Qiagen, SI04138407), siVKORC1(Qiagen, SI00760074), siFSP1(Qiagen, SI03082541), and siNC (Qiagen, SI03650318). The cells were transfected with 50 nM siRNAs using Lipofectamine® 3000 kit (Thermo Fisher Scientific) for 8 h at 37 °C in a 5% CO2 incubator. Following transfection, the culture medium was refreshed with complete medium, and the cells were reseeded in the complete culture medium for at least 8 h for subsequent experiments.
Statistical analysis
Each experiment was repeated at least three times unless otherwise stated. Statistics were display as the mean ± standard deviation (SD). Statistical analysis was performed using Student’s t test for comparisons between two groups or one-way analysis of variance (ANOVA) with post hoc Bonferroni correction for comparisons among >2 groups. A value of P < 0.05 was considered statistically significant.