Cell lines
GL261 was a kind gift from Professor Chunsheng Kang at Tianjin Medical University. CT2A was a kind gift from Professor Lei Zhang at Shaanxi Normal University. U-118 MG, U-87 MG, Vero and BHK-21 cells were purchased from ATCC. GL261-Luc and GL261-OVA cells were generated from GL261 cells via transduction of Luc-loaded lentivirus (Obio Technology Co. Ltd., Shanghai, China) and OVA-loaded lentivirus (Obio Technology Co. Ltd., Shanghai, China), respectively. The cells were grown in Eagle’s minimum essential medium (Corning, 10-010-CV) or Dulbecco’s modified Eagle’s medium (Gibco, 11965092) supplemented with 10% fetal bovine serum (FBS; 10099-141, Gibco, USA) and 1% penicillin/streptomycin (SV30010, HyClone). All the cells were cultured at 37 °C in 5% CO2.
Animals
The mice used in this study were 6- to 8-week-old females. C57BL/6J mice were purchased from Guangdong Medical Laboratory Animal Center. BALB/c-nu/nu mice were purchased from Guang Dong GemPharmatech Co., Ltd. C57BL/6JGpt-H11em1Cin (Tcra&Tcrb)/Gpt (OT-1) and B6/JGpt-Ighmem1Cd/Gpt (Ighm-KO) mice were purchased from GemPharmatech Co., Ltd. All the mice were bred in specific pathogen-free facilities. All the animal studies were approved by the Animal Ethical and Welfare Committee of Sun Yat-sen University.
OVM production and quantification
OVM-c6v1, OVM-GFP and OVM-iRFP were propagated in Vero cells and provided by Guangzhou Virotech Pharmaceutical Technology. The virus titer was determined via a CCID50 assay in BHK-21 cells.
Cell viability assay
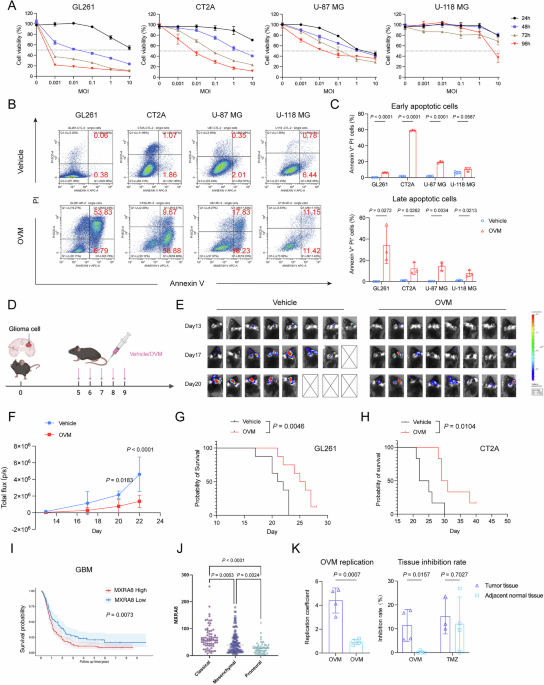
Glioma cells were seeded into 48-well plates at a density of 104 cells per well. Following overnight adherence, the cells were treated with OVM at MOIs of 0.001, 0.01, 0.1, 1, and 10. The samples were harvested at 24, 48, 72, and 96 h postinfection for analysis. The medium was aspirated and replaced with 200 µL of 3-(4,5-dimethylthiazol-2-yl)2,5-diphenyltetrazolium bromide (MTT; MP Biomedicals, 102227), 0.5 mg/mL final concentration, per well. After 2–4 h of incubation at 37 °C, the supernatant was carefully removed, and the MTT formazan precipitate was dissolved in 200 µL of dimethyl sulfoxide (DMSO; Sigma-Aldrich, 67-68-5). The absorbance was measured at 490 nm via a microplate reader (Synergy H1, BioTek).
ATP assay
Cell culture supernatants were obtained and centrifuged at 1000 × g for 10 min at 4 °C to remove cell debris. The concentrations of ATP were measured with ELISA kits (Beyotime Biotechnology, S0027) in accordance with the manufacturer’s instructions.
Tumor processing and ex vivo functional testing
This study was approved by the Institutional Review Board of Sun Yat-Sen University Cancer Center. GBM samples were freshly acquired during surgical resection. Live tumor tissues were minced into appropriately sized microtissue fragments (approximately 1 mm³) and uniformly seeded into 24-well plates. The plates were then incubated in a CO2 incubator. After 24 h of culture, the samples were treated with OVM (150 µL, titer: 1.16 × 108 CCID50/mL), TMZ (50 mg/mL) or vehicle control for 72 h. Subsequently, MTT stock solution was added to each well at a volume equivalent to 10% of the total well volume. After 2–4 h of incubation at 37 °C, the tissue fragments were analyzed via a tissue cell image analyzer. The analyzer captured both transmitted light images of the tissue fragments and quantified the area stained with formazan blue. The rate of drug inhibition was calculated via the following formula:
$${{{\rm{Inbibition}}}}\left( \% \right)=\left(1-\frac{{{{\rm{BA}}}}-{{{\rm{treated}}}}{{{\rm{sample}}}}/{{{\rm{A}}}}-{{{\rm{treatedsample}}}}}{{{{\rm{BA}}}}-{{{\rm{control}}}}{{{\rm{sample}}}}/{{{\rm{A}}}}-{{{\rm{controlsample}}}}}\right)\times 100 \%$$
(A: Total area of tissue fragments in transmitted light images prior to drug treatment. BA, blue area, area stained with formazan blue in transmitted light images after drug treatment).
qPCR
For the OVM replication assay, total RNA was extracted from tissue samples via TRIzol™ reagent (Thermo Fisher Scientific, 15596018CN). Viral RNA was quantified via one-step reverse transcription quantitative PCR (RT‒qPCR) via the FastKing One Step RT‒qPCR Kit (Probe, Tiangen Biotech, FP314-01) on an Applied Biosystems 7500 Fast Real-Time PCR System. A TaqMan® primer-probe set targeting the gene encoding nonstructural protein 2 (NSP2) of the OVM virus (designed and synthesized by Guangzhou Virotech Pharmaceutical Technology, Guangdong, China) was employed. To account for the nonspecific adsorption of viral particles to tissues, equal amounts of inactivated OVM-c6v1 were added to parallel samples as background controls. Viral replication within each tissue type was expressed as a replication coefficient, which was calculated as the ratio of viral copy numbers detected under standard culture conditions to those detected under inactivated conditions:
$${{{\rm{Replication\; coefficient}}}}=\frac{{{{\rm{Viral\; copies}}}}({{{\rm{standard\; culture}}}})}{{{{\rm{Viral\; copies}}}}({{{\rm{inactivated\; culture}}}})}$$
A replication coefficient greater than 2 was considered indicative of productive viral replication in the tissue. The PCR primers used were as follows: OVM-NSP2 forward primer: 5′-GGGATTCACTACACCTGCTTAGAC-3′; OVM-NSP2 reverse primer: 5′-GCTGACTCTGTCTGCGTAACC-3′; and OVM-NSP2 probe: 5′-CTCTCATCAGCAGCGAGCCTCCT-3′.
Animal models
For tumor inoculation, cells from all the lines were harvested via 0.05% trypsin, washed sequentially with complete medium and phosphate-buffered saline (PBS), and centrifuged. The cell pellets were then resuspended in PBS for subcutaneous injection (200 µL injection volume) or in a 1:1 PBS/methylcellulose mixture for intracranial injection (5 µL injection volume). 1 × 105 GL261, 1 × 104 GL261-Luc, 1 × 104 CT2A, and 1 × 105 GL261-OVA were injected intracranially. A total of 1 × 106 U-87 MG were injected subcutaneously. At 5–9 days post-inoculation, the mice were intravenously administered 300 µL of OVM (2 × 10⁷ CCID50/mL) via the tail vein.
For the splenectomy model, the mice were anesthetized via an intraperitoneal injection of pentobarbital sodium (50 mg/kg, 0.75% solution). The surgical site over the spleen was shaved and disinfected with iodophor. After the skin and fascia were incised, the spleen was exposed. Splenic vessels were ligated, and the spleen was excised following transection of its vascular connections and surrounding tissues. The wound was sutured and disinfected, and the animals were monitored postoperatively until they fully recovered from anesthesia.
In vivo imaging
The mice received an intraperitoneal injection of D-Luciferin potassium salt (150 mg/kg) (Absin, abs42075819) for bioluminescence substrate delivery. Prior to imaging, anesthesia was induced via the intraperitoneal administration of pentobarbital sodium (50 mg/kg, 0.75% solution). Bioluminescent signals were captured via an IVIS Spectrum system (PerkinElmer). The tumor burden was quantified by measuring the total flux (photons/second) within standardized regions of interest.
Serum IL-10 and TGF-β1 detection
The concentrations of IL-10 and TGF-β1 in the serum of glioma-bearing mice were detected via ELISA, as previously described in the literature [50]. A Mouse IL-10 ELISA Kit (MULTI SCIENCES, EK210) and a Mouse TGF-β1 ELISA Kit (MULTI SCIENCES, EK981) were used for detection, following the manufacturers’ instructions.
In vivo antibody administration
For the B-cell depletion experiments, the mice received intraperitoneal injections of 500 µg of anti-mouse CD19 (clone 1D3; Selleck, A2149) 1 day prior to tumor inoculation. Additional intraperitoneal injections of 250 µg of anti-mouse CD19 were administered on day 2 and day 7 posttumor inoculation. For the anti-PD-1/anti-PD-L1 antibody administration experiments, the mice received intraperitoneal injections of 250 µg of anti-mouse PD-1 antibody (clone RMP1-14; Selleck, A2122) or 250 µg of the rat IgG2a isotype control-InVivo (clone 2A3; Selleck, A2123) on days 5, 7, and 9 after tumor inoculation. For the anti-PD-L1 treatment group, the mice were intraperitoneally injected with 250 µg of anti-mouse PD-L1 antibody (clone RMP1-14; Selleck, A2115) or 250 µg of the rat IgG2b isotype control-InVivo (clone LTF-2; Selleck, A2116) at the same time points (days 5, 7, and 9 posttumor inoculation).
B cells, MZ B cells, Bst2+ B cells, CD8+ T cells, and DC isolation
Single-cell suspensions from mouse spleens were generated by mechanical dissociation through 40-µm cell strainers (BIOFIL, CSS-010-040), followed by red blood cell lysis (TIANGEN, RT122-02).
For total B-cell isolation, the EasySep™ Mouse B-Cell Isolation Kit (STEMCELL Technologies, 19854) was used. The single-cell suspension was adjusted to 1 × 108 cells/mL (0.25–2 mL), and an FcR blocker (20 μL per mL of sample) was then added to the suspension. The isolation cocktail (50 μL per mL of sample) was subsequently added, and the mixture was incubated at room temperature for 10 min. RapidSpheres™ were vortexed for 30 s, after which they were added to the sample (50 μL per mL of sample), and the mixture was incubated at room temperature for 2.5 min. The sample was subsequently brought to 2.5 mL with the recommended medium, and the lidless tube was placed into the kit-matched magnet (EasySep™, Catalog #18000) and incubated at room temperature for 2.5 min. Finally, the magnet and tube were inverted together, and the enriched B-cell suspension was poured into a new tube. The purity of the isolated total B cells was verified via flow cytometry, and the isolation efficiency exceeded 96% (Supplementary Fig. S8A).
For MZ B-cell enrichment (EasySep™ Mouse FITC Positive Selection Kit II, STEMCELL Technologies, 17668), purified B cells were adjusted to 1 × 108 cells/mL (0.1–2.5 mL) and transferred to the required tube. The FcR blocker (10 μL/mL sample) and Cr2-FITC antibody (1 μg/mL sample) were sequentially added, and the mixture was incubated for 15 min at room temperature. The selection cocktail (100 μL/mL sample) was added for 15 min. RapidSpheres™ (30 s) were vortexed and added (50 μL/mL sample) for 10 min of incubation. The sample was brought to 2.5 mL, and then the lidless tube was placed in the magnet for 5 min. The supernatant was discarded by inverting the assembly, and this separation process was repeated twice. The retained cells were MZ B cells. The purity of the enriched MZ B cells was verified via flow cytometry, and the isolation efficiency exceeded 93% (Supplementary Fig. S8B).
For Bst2+ B-cell isolation, purified MZ B cells were cultured in vitro with OVM (MOI of 10) and IL-4 (20 ng/mL) for 48 h. After incubation, the cells were harvested, washed twice with cold PBS containing 2% FBS, and stained with Bst2-APC antibody (eBioscience, 17-3172-82) for 30 min at 4 °C in the dark. The cells were then sorted to isolate Bst2+ and Bst2− B-cell subsets via a Beckman mFlo flow cytometer.
For CD8⁺ T-cell isolation, the density of the splenic cell suspensions was adjusted to 1 × 108 cells/mL. Biotin-Ab mixture (20 μL/mL sample) was added, and the mixture was incubated at 4 °C for 10 min. Streptavidin beads (200 μL/mL sample, vortexed to resuspend prior to use) were then added, followed by incubation at 4 °C for 10 min. Sorting buffer (2.5 mL) was added to the lidless tube, which was placed in the magnet for 5 min. The tube and magnet were inverted together, and the collected cell suspensions were purified from CD8+ T cells. The purity of the isolated CD8+ T cells was verified via flow cytometry, and the isolation efficiency exceeded 92% (Supplementary Fig. S8C).
For DC isolation (EasySep™ Mouse CD11c Positive Selection Kit, STEMCELL Technologies, 18780), the splenic single-cell suspension was adjusted to 1 × 108 cells/mL (0.5–2 mL), and an FcR blocker (60 μL per mL of sample) was added to the suspension. A selection cocktail (50 μL per mL of sample mixed from equal volumes of Component A and Component B) was prepared, incubated at room temperature for 5 min, and then added to the sample for 5 min of room temperature incubation. RapidSpheres™ were vortexed for 30 s and added (40 μL per mL of sample), followed by 3 min of room temperature incubation. The sample was topped to 2.5 mL with the recommended medium, and the lidless tube was placed into a kit-matched magnet (EasySep™, Catalog #18000) for 3 min. The magnet and tube were inverted to discard the supernatant, and this separation process was repeated three more times (4 total 3-min separations). The retained cells were resuspended in the desired medium to yield purified CD11c+ cells. The purity of the isolated DCs was verified via flow cytometry, and the isolation efficiency exceeded 90% (Supplementary Fig. S8D).
In vitro coculture
B cells were isolated from the spleens of glioma-bearing mice treated with either vehicle or OVM, whereas CD8+ T cells were isolated from vehicle-treated glioma-bearing mice. B cells and CD8+ T cells were cocultured in RPMI-1640 medium supplemented with 10% FBS at B:T ratios of 1:1, 5:1, or 10:1.
For transwell assays, B cells and CD8+ T cells were cocultured at a B:T ratio of 5:1 under two conditions: direct cell‒cell contact or physical separation by a transwell insert (BIOFIL, TCS016006).
For cell contact visualization, B cells were labeled with Cy5 (MCE, HY-D0819), and CD8+ T cells were labeled with CFSE (Invitrogen, C34554) prior to coculture at a B:T ratio of 5:1. After 24 h, dynamic live-cell imaging was performed via a fully motorized fluorescence microscope equipped with a live-cell incubation system (Nikon ECLIPSE Ti2).
For inhibitor studies targeting the immunological synapse, proteasome, lysosome, or ER-associated degradation (ERAD) pathways, B cells were pretreated for 24 h with vehicle (DMSO), the ICAM1–LFA interaction inhibitor A-286982 (Selleck, S3408) (1 µM), the proteasome inhibitor lactacystin (MCE, HY-16594) (2.5 µg/mL), the lysosome inhibitor cathepsin inhibitor I (TargetMol, 225120–65–0) (10 µM), or the ERAD inhibitor Eeyarestatin I (TargetMol, 412960–54–4) (3 µM). The cells were then cocultured with CD8+ T cells at a 5:1 ratio.
For MHC-I blockade, B cells and CD8+ T cells were cocultured at a 5:1 ratio in the presence of InVivoMAb anti-mouse MHC Class I (H-2) (clone M1/42.3.9.8; Bioxcell, BE0077) (100 µg/mL).
For the OVA uptake assay, B cells were isolated from the spleens of GL261-OVA-bearing mice and coincubated with ovalbumin-Cy5 (MCE, HY-NP055) (10 µg/mL) for 24 h, after which OVA-Cy5 uptake by B cells was assessed via flow cytometry.
Flow cytometry
For apoptosis detection following OVM infection, glioma cells were harvested with EDTA-free trypsin and centrifuged at 300 × g for 5 min (4 °C), and the supernatant was discarded. The cells were washed once with precooled PBS, recentrifuged, and resuspended in 100 μL of 1× Annexin V Binding Buffer (from the Annexin V-APC/PI Apoptosis Kit, Elabscience, E-CK-A217) to a concentration of 1 × 106 cells/mL. Then, 2.5 μL of Annexin V-APC and 2.5 μL of PI reagent (provided in the kit) were added to the cell suspension, which was vortexed gently and incubated at room temperature in the dark for 20 min before being diluted with 400 μL of precooled 1× Annexin V binding buffer for immediate analysis.
For cell sample preparation, single-cell suspensions from tumor tissues were prepared via the Tumor Dissociation Kit (Miltenyi Biotec, 130-096-730) according to the manufacturer’s protocol. Splenocytes were isolated as previously described, and single-cell suspensions derived from tumors, spleens, and PBMCs were subjected to red blood cell (RBC) lysis buffer treatment to remove erythrocytes.
Surface marker staining was performed as follows: first, the cells were stained with Fixable Viability Stain 780 (BD Biosciences, 565388) at a 1:1000 dilution (0.1 μL per 100 μL of cell suspension) with immediate vortexing and incubated for 15 min at room temperature in the dark to exclude dead cells. The cells were subsequently incubated with a CD16/CD32 monoclonal antibody (eBioscience, 14-0161-81) at a final concentration of 0.5 μg per test to block nonspecific Fc-mediated binding. Finally, the cells were stained with a panel of fluorochrome-conjugated surface antibodies and isotype controls at the manufacturer-recommended concentrations, with detailed information on all the antibodies and isotype controls provided in Supplementary Table 1. Staining was performed at 4 °C in the dark for 30 min, followed by two washes with precooled PBS.
For OVA-specific T-cell detection, after surface marker staining, the cells were resuspended in 50 μL of precooled PBS, and 1 μL of SIINFEKL-MHC tetramer-PE (BetterGen, BTG14028) was added to the 50 μL staining system. The mixture was incubated at 4 °C for 1 h in the dark and then washed twice with precooled PBS.
Intracellular staining was conducted via the Intracellular Fixation & Permeabilization Buffer Set (eBioscience, 88-8824-00): following surface staining and the final PBS wash, the supernatant was discarded, and the cell pellet was vortexed briefly to completely dissociate the cell clumps, with approximately 100 μL of residual volume retained. One hundred microliters of Intracellular Fixation Buffer was added to the cell pellet, followed by brief vortexing to mix thoroughly, and the mixture was incubated at room temperature for 30 min in the dark. After fixation, 2 mL of 1× permeabilization buffer was added, and the cells were centrifuged at 500 × g for 5 min at room temperature before the supernatant was discarded; this wash step was repeated once. The cell pellet was resuspended in 100 μL of 1× permeabilization buffer, and intracellular antibodies were added, followed by incubation at room temperature for 30 min in the dark. After staining, 2 mL of 1× permeabilization buffer was added, and the cells were centrifuged at 500 × g for 5 min at room temperature, after which the supernatant was discarded.
For the T-cell proliferation assay, CD8+ T cells were stained with 1 μM CFSE (Invitrogen, C34554) or 1 μM CellTrace Violet (Invitrogen, C34571) according to the manufacturer’s protocol: the cells were incubated for 20 min at room temperature or 37 °C protected from light, five times the original staining volume of culture medium (containing at least 1% protein) was added to quench free dye, and the mixture was incubated for an additional 5 min. The cells were pelleted by centrifugation and resuspended in fresh prewarmed complete culture medium, and the isolated B cells and labeled CD8+ T cells (including OT-1 CD8+ T cells) were cocultured at a 5:1 ratio (B:CD8+ T) in 48-well plates with positive control plates precoated with 3 µg/mL anti-CD3 (clone 145-2C11; BioLegend) and 5 µg/mL anti-CD28 (clone 37.51; BioLegend) antibodies.
For the T-cell function assay, positive control wells were treated with PMA (MCE, HY-18739) and ionomycin (MCE, HY-13434), and all experimental groups were supplemented with β-mercaptoethanol and brefeldin A (MCE, HY-16592). The cells were subjected to surface marker staining followed by the intracellular staining protocol described above.
All flow cytometric analyses were performed on a CytoFLEX flow cytometer (Beckman Coulter, USA). The data were analyzed with CytExpert software (Version 2.4.0.28), and histograms were specifically analyzed via FlowJo software (FlowJo, USA; Version 10.8.1). The detailed gating strategies are provided in Supplementary Fig. S9.
Adoptive transfer experiments
Splenic B cells were isolated from GL261-OVA-bearing C57BL/6J wild-type mice to obtain vehicle-treated B cells. MZ B cells were further purified from the spleens of GL261-OVA-bearing C57BL/6J wild-type mice and treated with OVM in vitro to generate Bst2+ and Bst2− B-cell subsets. Orthotopic GL261-OVA models were established in Ighm-KO mice, and 5 days later, the mice were adoptively transferred with vehicle B cells, Bst2+ B cells, Bst2− B cells, or vehicle control.
Multiplex immunofluorescence
Spleens were fixed in 4% paraformaldehyde (PFA), embedded in paraffin, and sectioned. The paraffin-embedded sections were incubated at 60 °C for 1 h, followed by deparaffinization and antigen retrieval. For antibody labeling, 0.5 µg of primary antibody was mixed with 1 µL of FlexLinker, and the volume was adjusted to 8 µL with FlexBuffer. The mixture was gently mixed and incubated for 5 min at room temperature in the dark. Subsequently, 2 µL of FlexQuencher was added, followed by gentle mixing and incubation for an additional 5 min under the same conditions. The final volume was adjusted to 50–100 µL with TBS. A mixture of multiple FlexAble-labeled antibodies was applied to the sections, which were subsequently incubated overnight at 4 °C in the dark. After incubation, the sections were mounted with Antifade Mounting Medium with DAPI (Beyotime Biotechnology, P0131) and scanned via a Digital Pathology Slide Scanner (KFBIO, KF-PRO-020). This multiplex immunofluorescence approach allows simultaneous detection of multiple antigens within the same tissue section by conjugating distinct fluorophores to different antibodies, enabling spatial mapping of cell populations and protein expression. The antibodies and reagents used were as follows: CD21/CD35 recombinant antibody (Proteintech, 84565-2-RR), CD19 polyclonal antibody (Proteintech, 27949-1-AP), BST2 polyclonal antibody (Proteintech, 30118-1-AP), FlexAble 2.0 Cora Lite® Plus 488 Antibody Labeling Kit for Rabbit IgG (Proteintech, KFA501), FlexAble 2.0 Cora Lite® Plus 594 Antibody Labeling Kit for Rabbit IgG (Proteintech, KFA509), and FlexAble 2.0 Cora Lite® Plus 555 Antibody Labeling Kit for Rabbit IgG (Proteintech, KFA502).
Single-cell RNA sequencing
Splenocytes were harvested from glioma-bearing mice receiving either vehicle or OVM treatment (n = 3 per group). Single-cell suspensions were assessed for quality and counted, with only those exhibiting >80% viability retained. The cells were washed, resuspended, adjusted to a final concentration of 700–1200 cells/µL, and then submitted to Majorbio Biopharm Technology Co., Ltd. (Shanghai, China) for single-cell RNA sequencing. Single-cell capture and barcoding were performed on the 10x Genomics Chromium platform, in which gel beads coated with unique molecular identifiers (UMIs) and cell barcodes were loaded at near saturation to ensure one bead per gel bead-in-emulsion (GEM). Within each GEM, the cells were lysed, and the mRNA was reverse transcribed to capture transcript information with cell-specific barcodes. Following GEM generation, the emulsion was broken, and the cDNA was purified, enriched, and amplified to construct 3′ gene expression libraries according to the manufacturer’s protocol (10x Genomics). Library sequencing was conducted on the Illumina XPlus platform, with initial sequencing and bioinformatic processing performed on Majorbio Co., Ltd.’s proprietary analysis platform (Shanghai, China).
The raw sequencing data were processed via CellRanger (v7.0.1), with the Mus musculus genome assembly GRCm38 (mm10) used as the reference. Downstream analysis was performed via the Seurat (v4.1.1) R package. To ensure high-quality data, we applied stringent quality control (QC) criteria: cells were retained only if they contained >200 detected genes, a mitochondrial gene percentage <10%, and a log10(genes)/log10(UMI) ratio >0.8. To exclude potential doublets, the scDblFinder (v1.11.4) package was employed to identify and remove multiplet droplets. After QC, the gene expression matrices were normalized via the normalizeData function, and the top 2000 highly variable features were identified.
To integrate data from different samples and minimize batch effects, we utilized the Harmony algorithm. Principal component analysis (PCA) was first performed on the scaled data (top 50 PCs). Unsupervised clustering was then conducted via a graph-based approach: a shared nearest neighbor (SNN) graph was constructed with k = 50 neighbors, followed by the Louvain algorithm for modularity optimization (via FindClusters with a resolution of 0.3). The resulting clusters were visualized via both t-distributed stochastic neighbor embedding (t-SNE) and uniform manifold approximation and projection (UMAP) based on Harmony-integrated embeddings (dims 1–50, seed = 1000).
Intercellular communication was inferred and quantified via CellChat (v1.6.1). We utilized the CellChatDB mouse database to analyze ligand‒receptor interactions. Separate CellChat objects were created for the “OVM” and “Vehicle” groups. The communication probabilities were calculated, and interactions were filtered if supported by fewer than 3 cells. The overall signaling network was aggregated, and network centrality scores were computed to identify key signaling hubs. Differential interaction analysis between groups was performed via the mergeCellChat and netVisual_diff interaction functions to visualize the strength and number of increased or decreased signaling pathways in response to OVM treatment.
To investigate the developmental transitions and lineage relationships (specifically for B cells), trajectory analysis was performed via Monocle3 (v1.3.1). The Seurat-processed data and UMAP embeddings were imported into the Monocle3 environment. A principal graph was learned across the cell manifold via the learn_graph function (with use_partition = FALSE). The cells were then ordered in pseudotime via the order_cells function to calculate their progression along the developmental path.
Statistical analysis
All the statistical analyses and graphical representations were performed via GraphPad Prism version 10.1.1, with the quantitative data conforming to normality (assessed via the Shapiro‒Wilk test) and homogeneity of variance (verified via the Brown‒Forsythe test) expressed as the mean ± standard deviation (SD); comparisons between two groups utilized unpaired two-tailed Student’s t tests, while multiple group comparisons employed one-way ANOVA followed by Dunnett’s post hoc test, and survival curves were analyzed via the log-rank (Mantel‒Cox) test. The 95% confidence intervals for all the statistical calculations are provided in Supplementary Table 2.