Reagents
TER was purchased from Targetmol (Boston, MA, USA). The PD-1/PD-L1 Blockade Bioassay kit was obtained from Promega (Madison, WI, USA). The cell counting kit-8 was purchased from Dojindo Molecular Technologies (Rockville, MD, USA). For cell dissociation and purification, a tumor dissociation kit, MACSmix tube rotator, gentleMACS C tubes, and gentleMACS dissociator were obtained from Miltenyi Biotec (Auburn, CA, USA). The anti-PD-L1 antibodies were purchased from BPS Bioscience (San Diego, CA, USA). Anti-PD-1 antibodies were purchased from Selleck Chemicals (Houston, TX, USA). A magnet for column-free cell separation, a Mouse CD8+ T Cell Isolation Kit, and polystyrene round-bottom tubes were purchased from STEMCELL Technologies (Vancouver, BC, Canada). Mouse IL-2 and IFN-γ ELISA kits were purchased from BD Biosciences (San Diego, CA, USA), Mouse GrB and TNF-α ELISA kits were purchased from Thermo Fisher Scientific (Waltham, MA, USA), and a Mouse Prf1 ELISA kit was obtained from Novus Biologicals (Centennial, CO, USA). Anti-CD8 antibodies were obtained from Cell Signaling Technology (Danvers, MA, USA).
Cell culture
Humanized PD-L1-expressing MC38 cells, derived from C57BL/6 mouse CRC, were obtained from the Shanghai Model Organisms Center (Shanghai, China). The cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) containing 10% heat-inactivated fetal bovine serum (FBS) and antibiotics (100 units/ml penicillin).
Recombinant Jurkat-T cells expressing human PD-1 and an NFAT reporter gene (hPD-1/NFAT Jurkat-T cells, #60535) and recombinant CHO-K1 cells expressing human PD-L1 and a T-cell receptor (TCR) activator (hPD-L1/TCR CHO-K1 cells, #60536) were obtained from BPS Bioscience. Jurkat and CHO-K1 cells were infected with human PD-1- and PD-L1-expressing lentiviruses (GeneChem, Shanghai). The cells were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium containing 10% heat-inactivated FBS and antibiotics (100 units/ml penicillin).
Human CRC cells (DLD-1, HCT116, and RKO) were obtained from the Korean Cell Line Bank (Seoul, Korea) and cultured in DMEM containing 10% heat-inactivated FBS and antibiotics (100 units/ml penicillin). All cell culture solutions were obtained from Hyclone Laboratories Inc. (Chicago, IL, USA). The cells were cultured at 37 °C in a humidified 5% CO2 incubator.
Cell viability assay
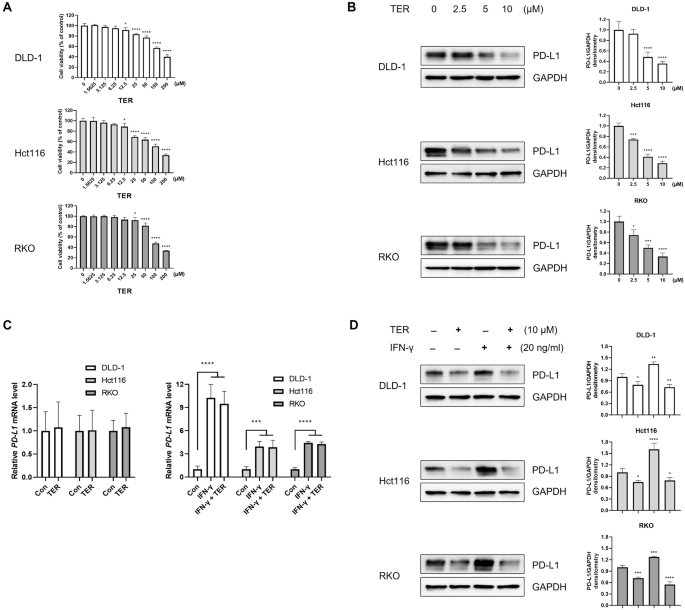
Cell viability was measured using the CCK assay (#CK04, Dojindo Molecular Technologies, Inc., Rockville, MD, USA) based on the manufacturer’s instructions. Briefly, cells were seeded into 96-well plates at a density of 1 × 104 cells per well and incubated overnight, followed by treatment with TER. A range of TER concentrations was added to the wells. After the designated incubation period, 10 μL of CCK solution was added to each well, and the plates were incubated for 2 h at 37 °C. The absorbance at 450 nm was measured using a microplate reader (Molecular Devices i3, San Jose, CA, USA), and cell viability was calculated.
Western blotting analysis
The cells were washed with 1× PBS, and total protein lysates were prepared using RIPA buffer containing 1% NP-40 lysis buffer and protease inhibitor cocktail tablets (Roche, Basel, Switzerland). Protein concentration was measured using the Bio-Rad protein assay. Equivalent amounts of protein were separated by 8–15% SDS-PAGE and transferred to nitrocellulose membranes (GE Healthcare, Munich, Germany). The membranes were blocked with 5% nonfat dry milk at room temperature (20–25 °C) for 1 h and incubated overnight at 4 °C with primary antibodies. The membranes were washed three times with 1× Tris-buffered saline for 10 min per wash, followed by incubation with the corresponding secondary antibody. After washing, protein detection was performed using the ChemiDoc Touch Imaging System (Bio-Rad, Hercules, USA). GAPDH was used as a loading control for protein normalization. The original uncropped images for all western blots are provided in the Supplementary Information (Supplementary Figs. S1–S7).
Quantitative reverse transcription–polymerase chain reaction (qRT–PCR)
Total RNA was isolated using the RiboEx Total RNA Extraction Kit (GeneAll Biotechnology, Seoul, Korea). Complementary DNA (cDNA) was synthesized using the AccuPower CycleScript RT premix (dT20) (Bioneer, Daejeon, Korea). For the reverse transcription reaction, 1 µg of total RNA was used in a final cDNA reaction volume of 20 µL. Quantitative PCR was performed using a QuantStudio 3D Digital PCR System (Thermo Fisher Scientific, Waltham, MA, USA) along with the AccuPower 2X Greenstar qPCR master mix (Bioneer). The protocol consisted of 40 cycles of denaturation at 95 °C for 15 s, followed by annealing and extension at 60 °C for 30 s per cycle. All reagents were used according to the manufacturer’s instructions. The relative expression of target mRNA was normalized to GAPDH expression as an endogenous control.
Competitive ELISA
A competitive ELISA was performed using the PD-1/PD-L1 inhibitor ELISA screening kit (BPS Bioscience, #72005, San Diego, CA, USA), following the manufacturer’s instructions. The αPD-L1 antibody (#71213) served as the positive control. Recombinant hPD-L1 (#71104, BPS Bioscience) was coated onto 96-well plates (Corning Inc., New York, NY, USA) at a concentration of 1 mg/mL in PBS and incubated overnight. The plates were then washed with PBS containing 0.1% Tween (PBS-T), blocked with 2% BSA in PBS for 1 h at room temperature, and re-washed. Next, 5 μL of 0.5 mg/mL biotinylated hPD-1 (#71109, BPS Bioscience) was added, and the plates were incubated for 2 h at room temperature. After three washes in PBS-T, 50 μL of 0.2 mg/mL HRP-conjugated streptavidin was added and the plates were incubated for 1 h. Finally, the plates were washed three times with PBS-T, and chemiluminescence was measured using a SpectraMax L Luminometer (Molecular Devices, San Jose, CA, USA).
PD-1/PD-L1 blockade bioassay
To determine the effectiveness of TER in blocking the PD-1/PD-L1 interaction, a cell-based PD-1/PD-L1 blockade bioassay was performed using the PD-1/PD-L1 blockade bioassay kit (#J4011, Promega, Madison, WI, USA), with slight modifications to the manufacturer’s instructions. In this assay, 5 × 104 PD-L1 aAPC/CHO-K1 cells were seeded into 96-well plates containing DMEM supplemented with 10% FBS. On the experiment day, the medium was removed, and PD-L1-blocking antibody or TER (as indicated) was added along with 1 × 105 PD-1 effector cells. Luminescence was measured using the GloMax® Explorer Multimode Microplate Reader (Promega) after adding Bio-Glo™ Reagent (Promega, #G7940). The results are expressed as the mean ± standard error of the mean from four independent experiments.
Tumor-infiltrating T cells (TILs) isolation
A humanized PD-1 mouse model (C57BL/6J background, genetically engineered to express the full-length human PD-1 protein) was obtained from the Shanghai Model Organisms Center (Shanghai, China). The animal experiments were conducted in compliance with the guidelines established by the Institutional Animal Care and Use Committee (IACUC) of the Korea Institute of Oriental Medicine (KIOM) and approved by the IACUC (approval number: KIOM-D-23-099). To extract tumor-infiltrating lymphocytes (TILs, CD8+ T cells), MC38 tumor tissues were collected and digested with collagenase IV (0.5 mg/mL) at 37 °C for 1 h. The resulting cell suspension was strained twice using 100-μm and 40-μm cell strainers (SPL) to isolate single cells. TILs were isolated from the single-cell suspension using negative selection beads according to the manufacturer’s instructions (STEMCELL Technologies Inc., Vancouver, Canada).
Co-culture experiments
Tumor-infiltrating CD8+ T cells (1 × 106 cells) were used as effector cells and activated with Dynabeads T Activator CD3/CD28 (Life Technologies, Carlsbad, CA, USA) for 72 h at 37 °C. Murine hPD-L1 MC38 cells, which served as target cells, were labeled using the CellTrace™ Far Red Cell Proliferation Kit (Thermo Fisher Scientific, Waltham, MA, USA) and treated with IFN-γ (10 ng/mL) for 24 h at 37 °C to induce PD-L1 expression. hPD-L1 MC38 cells (5 × 104 cells) were co-cultured with the activated CD8+ T cells (2.5 × 105 cells) at an effector-to-target cell ratio of 5:1 for 72 h at 37 °C in the presence of different concentrations of TER. Then, the plates were washed with PBS, and the remaining viable cancer cells were stained with crystal violet solution and quantified using a SpectraMax i3 microplate reader at 540 nm.
In vivo analysis
The experimental mice (male, n = 6) were housed in a specific pathogen-free facility and anesthetized using a 2% isoflurane–oxygen mixture administered through an Avesko inhalation system. To induce subcutaneous tumor formation, hPD-L1 MC38 cells were injected into the right flank of humanized PD-1 mice. Each mouse received 200 μL of cell suspension (5 × 105 cells/mouse). Tumor progression was assessed biweekly using a digital caliper (Hi-Tech Diamond, Westmont, IL, USA). Tumor volume was calculated using the formula: (W2 × L)/2, where W represents the short diameter and L represents the long diameter.
Once the tumors reached a volume of 100 mm3, the mice were divided into several experimental groups. The control group received vehicle [PBS, 10 mL/kg, orally (q.d., i.g.)], whereas the other groups were treated with either low or high doses of TER [10 or 30 mg/kg (mpk), administered intraperitoneally (q.d., i.p.)], or a CD8 depletion antibody (#BP0061, Bio Cell, West Lebanon, NH, USA). TER was administered continuously for 16 consecutive days. In contrast, the CD8 depletion antibody was administered twice a week through intragastric injections beginning after tumor implantation. On day 16 following treatment, the animals were euthanized for further analysis.
Tissue dissociation for cell isolation and flow cytometry
Tumor tissues were collected in RPMI Medium 1640 containing 10% heat-inactivated FBS, 1% penicillin and streptomycin (all from HyClone™), minced, and incubated for 30 min in DPBS (Well-gene) with 0.5 mg/mL Collagenase IV (Sigma) at 37 °C. The tissues were squeezed through a 100 μm cell strainer (#93100, SPL), re-filtered through a 70 μm cell strainer (#93070, SPL), and incubated for 5 min in Red Blood Cell Lysis Buffer (#10-548E, Lonza). Flow cytometry was conducted to determine cell surface antigen expression by a 30-min incubation on ice with the following antibodies: mouse-specific monoclonal antibodies PerCP/Cyanine5.5 anti-mouse CD45 (#103131, BioLegend), APC anti-mouse CD8a (#100712, BioLegend), corresponding isotype control mAbs PerCP/Cy5.5 Rat IgG2b (#400632, BioLegend), and APC Rat IgG2a (#400512, BioLegend). The cells were washed three times with PBS containing 2% FBS, fixed in suspension with 4% paraformaldehyde, and stored at 4 °C until subsequent analysis with a CytoFLEX flow cell counter (Beckman). The data were analyzed using Kaluza analysis software (Beckman Coulter).
Surface plasmon resonance analysis
SPR was used to measure binding affinity between PD-L1 and PD-1 proteins using a Biacore T200 biosensor (GE HealthCare Technologies, Inc., Chicago, IL, USA) with a nitrilotriacetic acid (NTA) sensor chip (GE HealthCare). The NTA surfaces were washed with 350 mM EDTA and loaded with 0.5 mM NiCl2. The hPD-1 protein (#10377-H08H, Sino Biologicals) was bound to the NTA sensor chip using an affinity capture method. Various concentrations of analytes were prepared by diluting in HBS-P buffer (10 mM HEPES, pH 7.4, 150 mM NaCl, and 0.005% surfactant P20). The equilibrium dissociation constants (KD; kd/ka) were calculated using sensorgrams.
Molecular docking analysis
To examine the interaction between human PD-1 and TER, molecular docking was conducted using the CB-Dock2 platform, an automated docking tool capable of cavity detection and AutoDock Vina-based docking [46]. The target protein structure was obtained from the Protein Data Bank (PDB ID: 3RRQ) [47] and used in its native form without any mutations or modifications. TER was used as a ligand, and its 3D structure file (.sdf) was downloaded from the PubChem database (CID: 54684141) [48]. CB-Dock2 automatically detected potential binding cavities within the PD-1 structure, and the ligand was docked using AutoDock Vina. Subsequently, the interaction between PD-1 and Teriflunomide was visualized with the built-in visualization tools of the CB-Dock2 platform.
Statistical analysis
The data are presented as ratios relative to their respective control values. The results are shown as means ± standard error of the mean (SEM) based on three independent experimental replicates. Differences between the mean values within each group were evaluated using a Student’s t test, whereas comparisons across multiple groups were done using a one-way analysis of variance, followed by Tukey’s post hoc test. The statistical analyses were performed using GraphPad Prism version 8.0 (GraphPad Software, San Diego, CA, USA).
Ethical approval and consent to participate
All animal experiments were performed in accordance with the relevant guidelines and regulations. The study was approved by the Institutional Animal Care and Use Committee (IACUC) of the Korea Institute of Oriental Medicine (KIOM) (approval number: KIOM-D-23-099).