In silico analysis
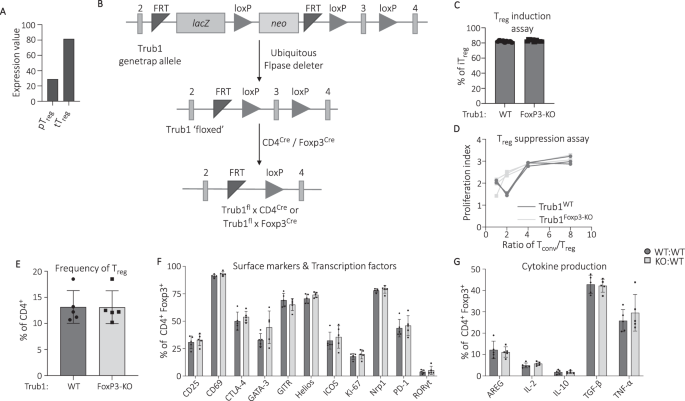
Data on Trub1 mRNA expression across different immune cell subsets were obtained as expression values from Gene Skyline hosted by immgen.org.
Animals
Congenic CD45.1 mice (B6.SJL-Ptprca.Pepcb/BoyJ) were purchased from Jackson Laboratories (strain 002014) and crossed to Foxp3Thy1.1 reporter mice [45]. Trub1 genetrap heterozygous frozen embryos (C57BL/6N-Trub1tm1a(EUCOMM)Hmgu) were purchased from the MMRRC repository (UC Davis, University of California, CA, USA), then bred to mice with constitutive expression of flippase (B6.SJL-Tg(ACTFLPe)9205Dym/J, strain 005703, Jackson Laboratories) to create the conditional Trub1 allele (short Trub1flox). Trub1flox/+ mice were bred to CD4Cre transgenic mice for T cell-specific Trub1 deficiency (Tg(Cd4-cre)1Cwi [46], C57BL/6J background, Trub1Tcell-KO for short) or Foxp3CreYFP knockin mice for Treg-specific Trub1 deficiency (C57BL/6.129(Cg)-Foxp3tm4(Cre)Ayr/J [47], Trub1Foxp3-KO for short). Of note, the Foxp3YFPCre knock-in allele has been described to display promiscuous Cre recombinase activity for some conditional alleles [48]. In our mouse colony, we identified germline Cre recombinase activity resulting in mice with global Trub1 deficiency (Trub1All-KO for short).
Ethical approval
All methods were performed in accordance with the relevant guidelines and regulations. Specifically, sample sizes for mouse experiments were chosen based on power calculations and pilot data, in conjunction with the Animal Ethics Committee, to allow for robust sensitivity without excessive animal use. Mice were used at 8-12 weeks of age. Both male and female mice were used in the experiments. Mice were housed in specific pathogen-free conditions under a 12-h light/dark cycle in a temperature- and humidity-controlled room. Age- and sex-matched animals were used for each experiment. Animals were also co-housed when possible. No other randomization procedures were used. All animal procedures were approved by the KU Leuven Animal Ethics Committee (approvals: 150/2019, 008/2019, 228/2024, 74/2018, 060/2024).
Generation of mixed bone marrow chimera mice
Bone marrow was extracted from CD45.2 Trub1CD4/Foxp3/ALL-KO, Trub1WT or CD45.1 wild-type mice by crushing the femurs with a mortar and pestle. Cells were then resuspended in Tris-NH4Cl red blood cell (RBC) lysis buffer to lyse the RBCs. Mixed bone marrow chimeric mice were generated by co-transplanting an equal mixture of bone marrow from CD45.2 Trub1CD4/Foxp3/ALL-KO mice with CD45.1 wild-type mice (KO: WT). To control for strain differences, mice were also co-transplanted with an equal mixture of bone marrow from CD45.2 Trub1WT and CD45.1 wild-type mice (WT: WT). A total of 20 × 106 cells were injected intravenously into lethally irradiated mice (9.5Gy gamma-irradiation). Chimeric mice were analysed 8-10 weeks post-transplantation.
RNA isolation, reverse transcription and quantitative PCR
Total RNA was isolated from spleens and lymph nodes from Trub1-deficient mice and wild-type C57BL/6J mice using TRIzol reagent (Invitrogen). 1 μg of RNA was reverse-transcribed using Maxima H Minus First Strand cDNA Synthesis Kit (Thermo Scientific). Real-time qPCR was performed using Fast SYBR Green Master Mix (Applied Biosystems) with a StepOnePlus Real-Time PCR System (Applied Biosystems), according to the manufacturer’s instructions. The β-actin transcript was used as an internal control to normalise the transcript levels of Trub1. The following primers were used for qPCR reactions: Trub1-forward: 5’-TGAACCGGCTGAAGGAGAAG-3’, Trub1-reverse: 5’-CCAACCACCAGAACACCTTG-3’, β-actin-forward: 5’-CATCACTATTGGCAACGAGC-3’, β-actin-reverse: 5’-ACGCAGCTCAGTAACAGTCC-3’. Technical duplicates, as well as Reverse Transcriptase minus and no template control reactions, were performed alongside (data not shown). Relative quantification of Trub1 expression was evaluated using the ΔΔCt method [49]. The fold-change in the relative gene expression was determined by calculating the 2−ΔΔCt value.
Flow cytometry and cell sorting
Antibodies and fixable viability dye were purchased from eBioscience, Biolegend or BD Biosciences as indicated below.
Splenocytes were resuspended with Tris-NH4Cl red blood cell (RBC) lysis buffer to lyse the RBCs. Cells were then washed twice with PBS, stained with fixable viability dye (eBioscience) and, where applicable, stained for anti-CD69 (H1.2F3, Biolegend) at 4 °C for 20 min prior to fixation. Cells were then fixed using eBioscience™ Foxp3/Transcription Factor fixation/permeabilization kit (#00-5521-00, eBioscience) according to the manufacturer’s instructions. Cells were stained overnight at 4 °C with the panels covering: anti-CD45 (30-F11), anti-CD3 (145-2C11, 17A2,), anti-CD19 (B4), anti-CD11c (N418), anti-MHCII (M5/114.15.2), anti-CD172a (P84), anti-CD64 (X54-5/7.1), anti-PDCA-1 (927), anti-XCR-1 (ZET), anti-Ly6G (1A8), anti-CD44 (IM7), anti-ICOS (C398.4 A), anti-PD-1 (29 F.1A12), anti-KLRG1 (2F1/KLRG1), anti-IL-2 (JES6-5H4), anti-IL-4 (11B11), anti-IL-22 (Poly5164), anti-TGF-β (TW7-15B4), anti-TNF-α (MP6-XT22), anti-CD127 (S18006K), anti-Neuropilin-1 (3E12), anti-FoxP3 (FKJ-16s) (all from Biolegend); anti-CD3 (KT3), anti-CD19 (B4,6D5), anti-CD4 (RM4-5) (all from Bio-Rad); anti-CD4 (GK1.5), anti-CD103 (M290), anti-Ly6C (AL-21, anti-Siglec-F (E50-2440), anti-CD117 (c-kit) (2B8), anti-CD8α (53-6.7), anti-IgE (R35-72), anti-CD11b (M1/70), anti-CD95 (Jo2), anti-TCR-β (H57-597), anti-CD62L (MEL-14), anti-CTLA4 (UC10-4F10-11), anti-CD127 (SB/199), anti-GATA-3 (L50-823), anti-CD25 (PC61), anti-T-bet (4B10), anti-GITR (DTA-1), anti-IL-17 (TC11-18H10), anti-IL-6 (MP5-20F3), anti-IL11β (B122), anti-IFN-γ (XMG1.2), anti-RORγt (Q31-378) (all from BD Biosciences); anti-IgM (II/41), anti-NK1.1 (PK126), anti-Eomes (Dan11mag), anti-Neuropilin-1 (3DS304M), anti-ST2 (RMST2-2), anti-IL-10 (JES5-16A3), anti-IL-3 (MP2-8F8), anti-IL-13 (eBio13A), anti-IL-21 (FFA21), anti-RORγt (B2D), anti-Ki-67 (SolA15), anti-Helios (22F6), anti-GATA-3 (TWAJ), anti-IL11β (NJTEN3), anti-Ki-67 (16A8, SolA15) (all from eBioscience); anti-FoxP3 (REA788), anti-Helios (REA829) (both from Miltenyi Biotec); anti-Amphiregulin (polyclonal), anti-IL-5 (TRFK5) (both from R&D Systems).
To assess cytokine production, cells were stimulated in the presence of phorbol myristate acetate (50 ng/mL), ionomycin (1 µg/mL) and brefeldin A (2 µg/mL) (all Tocris Biotechne) for 4 h at 37 °C in RPMI supplemented with 10% of fetal bovine serum, 0.1 mM of non-essential amino acid, 50 UI/mL of penicillin G, 5 μg/mL streptomycin, 1 mM sodium pyruvate, 10 μg/mL β-mercaptoethanol (all from GIBCO, Thermo Fisher Scientific). Cells were then stained with fixable viability dye, followed by fixation with 2% Formaldehyde (VWR chemicals, Radnor, Pennsylvania, PA, USA) in PBS at room temperature for 30 min and permeabilized with 1x eBioscience permeabilization buffer (#00-8333-56, eBioscience) according to the manufacturer’s instructions prior to antibody staining.
Data were acquired on the BD FACSymphony A5 (BD Biosciences) and Aurora spectral analyser (Cytek). Data was analysed using FlowJo (v10.9.0, BD Biosciences) and FCS Express (v7.28.0019, De Novo Software). Median Fluorescence Intensity (MFI) is reported on marker-positive populations exclusively. Samples were excluded when the number of cells in the parent population was less than 100 or when the frequency of positive cells was less than 0.01%. In these cases, figure legends indicate where populations were removed from data presentation.
For cell sorting, splenic CD4 T cells were purified using the MojoSort™ Mouse CD4 T Cell Isolation Kit (Biolegend) according to the manufacturer’s instructions, followed by surface staining with anti-CD4 (GK1.5, BD Biosciences), anti-CD62L (MEL-14, BD Biosciences), anti-CD44 (IM7, Biolegend) and anti-CD90.1/Thy1.1 (HIS51, eBioscience) (when using congenic Foxp3Thy1.1 reporter mice) or CD25 (PC61, eBioscience) when using CD45.2 mice carrying conditional or knockout Trub1 alleles. DAPI (Thermo Scientific) was added for dead cell exclusion. Naïve CD4 T cells (gated as Thy1.1neg or CD25neg non-Treg) were defined as CD4+ CD44neg CD62L+ Thy1.1neg (or CD25neg). Treg were defined as CD4+ Thy1.1neg (or CD25neg) and post-sort stained for Foxp3 for additional purity confirmation. Cell sorting was performed on a BD FACSAria III instrument (BD Biosciences).
In vitro polarisation assay
96-well plates were pre-coated either overnight at 4 °C or 1 h at 37 °C with 5 µg/mL anti-CD3ε (145-2C11, Invitrogen) and 5 µg/mL anti-CD28 (37.51, Invitrogen). After washing the wells with PBS, 2 × 105 sort-purified naïve CD4 T cells were added and activated for 5 days in RPMI 1640 medium (Lonza) supplemented with 10% of fetal bovine serum, 0.1 mM of non-essential amino acid, 50 UI/mL of penicillin G, 5 μg/mL streptomycin, 1 mM sodium pyruvate, 10 μg/mL β-mercaptoethanol (all from GIBCO, Thermo Fisher Scientific). The following cytokine/neutralizing antibody cocktails were added to induce T helper cell polarization: TH1: IL-12p70 (10 ng/mL), IL-2 (1 ng/mL); TH2: anti-IFN-γ (10 μg/mL, XMG1.2, Invitrogen), IL-4 (10 ng/mL), IL-2 (1 ng/mL); TH17: anti-IFN-γ (10 μg/mL, XMG1.2, Invitrogen), IL-6 (40 ng/mL), TGF-β (1 ng/mL); Treg: IL-2 (20 ng/mL), TGF-β (5 ng/mL), anti-IFN-γ (10 µg/mL, XMG1.2, Invitrogen).
In vitro suppression of conventional T cells by Treg
Tregs from Trub1Foxp3-KO or from wild-type littermates were sort-purified as stated above. Conventional T cells (Tconv, CD4+ Thy1.1neg) were sort-purified from spleens of congenic CD45.1 Foxp3Thy1.1 reporter mice and were stained with CellTraceTM Violet (CTV, Thermo Fisher Scientific) as per the manufacturer’s protocol. 1 × 105 labelled Tconv were cultured in a U-bottom 96-well with CTV-unlabelled Tregs at different Treg: Tconv ratios (1:1, 1:2, 1:4, 1:8) in the presence of anti-CD3ε (1 μg/mL, 145-2C11, Invitrogen) and splenocytes from Rag1–/– mice (5 × 104 cells). After five days of co-culture, the cells were stained with fixable viability dye (eBioscience) and anti-CD45.1 (A20, eBioscience) and data were acquired on a BD FACSymphony A5 (BD Biosciences). Proliferation of CD45.1 Tconv was evaluated via dilution of CTV and labelled Tconv cultured for 5 days in the absence of Tregs were used as a control for proliferation.
MC38 tumour mouse model
The colon adenocarcinoma MC38 cell line was kindly provided by Prof. Gabriele Bergers, VIB-KU Leuven, Belgium and maintained in DMEM (Gibco) supplemented with 10% fetal bovine serum (Thermo Fisher), 1% glutamine, 100 U/mL penicillin and 100 U/mL streptomycin (All Gibco). 5 × 105 cells were injected subcutaneously in the left and right flanks of Trub1ALL-KO or wild-type littermates. Tumour width and length were followed at day 10, 12, 14 and 17 post-injection and tumour volume was calculated as: (width)2 x length x 0.52. For the measurement of tumour width and length, the operator was blinded to the experiment groups and the genotype of the animals. Mice were sacrificed 17 days after injection and for flow cytometry analysis, tumours, inguinal (tumour-draining), mesenteric, brachial and axillary lymph nodes (non-tumour draining) were isolated. Leukocytes from the lymph nodes were isolated via mechanical dissociation using frosted glass slides and were filtered through a 70 μm mesh. To obtain tumour-infiltrating leukocytes, tumours from both the left and right flanks were pooled and finely chopped. The pieces were transferred to a gentleMACSTM C Tube (Miltenyi Biotec) containing the enzyme mix with enzymes A, D and R (Tumour Dissociation Kit, Miltenyi Biotec). The enzyme mix was prepared as per the manufacturer’s instructions. The tissue was dissociated using the soft/medium program on a gentleMACSTM Octo Dissociator (Miltenyi Biotec). The single cell suspension was then filtered through a 70 μm mesh and RBC lysis was performed as described above. Tumour-infiltrating lymphocytes were further enriched via density gradient centrifugation using 40% v/v and 80% v/v Percoll (Cytiva). Antibody staining for flow cytometry-based analysis of the leukocytes and for assessing the cytokine production was performed as described above.
Influenza mouse model
Trub1KO or Trub1Tcell-KO mice or wild-type littermates received intranasal instillation of 10 plaque-forming units (PFU) of mouse-adapted influenza virus A/H3N2/Ishikawa/7/82 or PBS (10 μl per nostril) under inhalation anaesthesia with isoflurane (1.5–2% in 100% oxygen, Piramal Critical Care). Body weight and clinical score based on body weight, respiratory parameters and condition (adapted from [50]) were followed daily for up to 7 days post-inoculation. For the body weight measurement and evaluation of clinical score, the operator was blinded to the experiment groups and the genotype of animals. Mice were sacrificed 7 days after influenza infection. Spleen and bronchoalveolar lavage (BAL) fluid were collected. BAL fluid was collected after three lung lavages with 700 μL 0.9% NaCl (B. Braun). Immunophenotyping was performed on the cells obtained from BAL fluid and spleen via high-parameter flow cytometry as detailed above.
Micro-computed tomography (μCT) for influenza-infected mice
A whole-body small animal μCT scanner (SkyScan 1278, Bruker micro-CT. Kontich, Belgium) was used to acquire lung μCT data with the following parameters: 50kVP X-ray source, 1 mm aluminium X-ray filter, 350 μA current, 150 ms exposure time per projection, 0.9° increments over a total angle of 220° (adapted from [50]). µCT data was reconstructed, visualised and segmented using NRecon, DataViewer and CTan software, all provided by the manufacturer. Non-aerated lung volume (NALV), which is the imaging-derived biomarker of pulmonary lesions, was quantified based on the manual delineation of a region of interest (ROI), which resulted in a volume of interest (VOI) covering the entire lung and the predetermined threshold of 108-225 greyscale (adapted from [50]). For the imaging and visualisation, the operator was blinded to the experiment groups and the genotype of the animals.
Statistics
Statistical analyses were performed using GraphPad Prism, version 9.0 (GraphPad Software, San Diego, CA, USA) with a non-parametric Mann–Whitney U test due to non-normal data distribution. For the statistical analysis of clinical score progression, body weight loss and non-aerated lung volume in the influenza model and for tumour volume progression in the MC38 tumour model, two-way ANOVA with Tukey’s multiple comparison was used. p < 0.05 was considered significant. When linear mixed-effects models (LMMs) were applied and significance was assessed using ANOVA with F-tests, this is mentioned in the respective figure legend.