Pizzi, M., Margolskee, E. & Inghirami, G. Pathogenesis of peripheral T cell lymphoma. Annu. Rev. Pathol. 13, 293–320 (2018).
Fiore, D. et al. Peripheral T cell lymphomas: from the bench to the clinic. Nat. Rev. Cancer 20, 323–342 (2020).
World Health Organization. Haematolymphoid Tumours 5th edn, Vol. 11 (WHO, 2024).
Cortés, J. R. & Palomero, T. Biology and molecular pathogenesis of mature T-cell lymphomas. Cold Spring Harb. Perspect. Med. 11, a035402 (2021).
de Leval, L. et al. The gene expression profile of nodal peripheral T-cell lymphoma demonstrates a molecular link between angioimmunoblastic T-cell lymphoma (AITL) and follicular helper T (TFH) cells. Blood 109, 4952–4963 (2007).
Bishop, D. C. et al. Development of CAR T-cell lymphoma in 2 of 10 patients effectively treated with piggyBac-modified CD19 CAR T cells. Blood 138, 1504–1509 (2021). This study, together with Harrison et al. (2023), Kobbe et al. (2024), Braun et al. (2025) and Perica et al. (2025), identified T cell lymphomas that originated from CAR T cells.
Harrison, S. J. et al. CAR plus T-cell lymphoma post ciltacabtagene autoleucel therapy for relapsed refractory multiple myeloma. Blood 142, 6939 (2023).
Harrison, S. J. et al. CAR+ T-cell lymphoma after cilta-cel therapy for relapsed or refractory myeloma. N. Engl. J. Med. 392, 677–685 (2025).
Kobbe, G. et al. Aggressive lymphoma after CD19 CAR T-cell therapy. N. Engl. J. Med. 391, 1217–1226 (2024).
Teras, L. R. et al. 2016 US lymphoid malignancy statistics by World Health Organization subtypes. CA Cancer J. Clin. 66, 443–459 (2016).
Liu, S. et al. Epidemiological characteristics of peripheral T-cell lymphoma: a population-based study. Front. Oncol. 12, 863269 (2022).
Dustin, M. L. The immunological synapse. Cancer Immunol. Res. 2, 1023–1033 (2014).
Rodilla-Ramirez, C. et al. T cell protrusions enable fast, localised initiation of CAR signalling. Preprint at bioRxiv https://doi.org/10.1101/2025.07.08.662959v1 (2025).
De Boer, R. J., Homann, D. & Perelson, A. S. Different dynamics of CD4+ and CD8+ T cell responses during and after acute lymphocytic choriomeningitis virus infection. J. Immunol. 171, 3928–3935 (2003).
Bretscher, P. A. A two-step, two-signal model for the primary activation of precursor helper T cells. Proc. Natl Acad. Sci. USA 96, 185–190 (1999).
Nishimura, H., Nose, M., Hiai, H., Minato, N. & Honjo, T. Development of lupus-like autoimmune diseases by disruption of the PD-1 gene encoding an ITIM motif-carrying immunoreceptor. Immunity 11, 141–151 (1999).
Curtsinger, J. M., Gerner, M. Y., Lins, D. C. & Mescher, M. F. Signal 3 availability limits the CD8 T cell response to a solid tumor. J. Immunol. 178, 6752–6760 (2007).
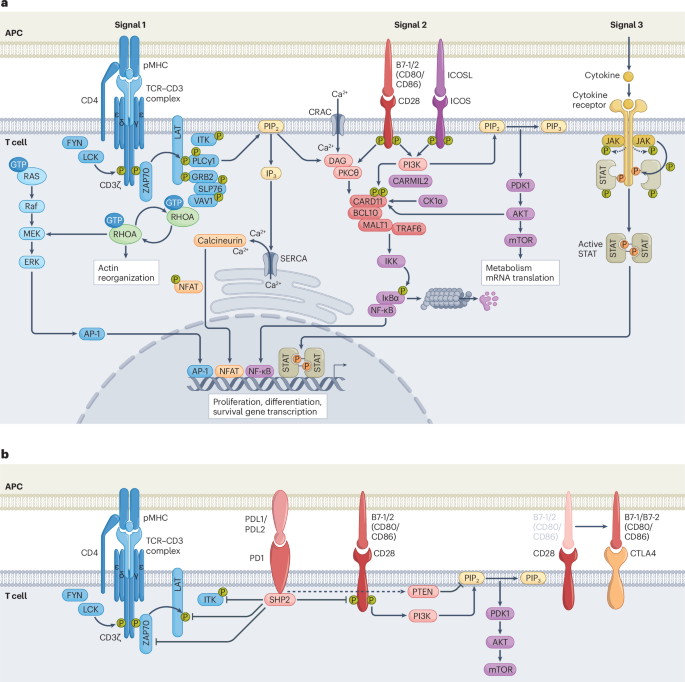
Gaud, G., Lesourne, R. & Love, P. E. Regulatory mechanisms in T cell receptor signalling. Nat. Rev. Immunol. 18, 485–497 (2018).
Shah, K., Al-Haidari, A., Sun, J. & Kazi, J. U. T cell receptor (TCR) signaling in health and disease. Signal. Transduct. Target. Ther. 6, 412 (2021).
Davis, S. J. & van der Merwe, P. A. The kinetic-segregation model: TCR triggering and beyond. Nat. Immunol. 7, 803–809 (2006).
Weiss, A. Peeking into the black box of T cell receptor signaling. Annu. Rev. Immunol. 42, 1–20 (2024).
Burkhardt, A. L. et al. Temporal regulation of non-transmembrane protein tyrosine kinase enzyme activity following T cell antigen receptor engagement. J. Biol. Chem. 269, 23642–23647 (1994).
Iwashima, M., Irving, B. A., van Oers, N. S., Chan, A. C. & Weiss, A. Sequential interactions of the TCR with two distinct cytoplasmic tyrosine kinases. Science 263, 1136–1139 (1994).
Chan, A. C., Iwashima, M., Turck, C. W. & Weiss, A. ZAP-70: a 70 kd protein-tyrosine kinase that associates with the TCR zeta chain. Cell 71, 649–662 (1992).
Balagopalan, L., Coussens, N. P., Sherman, E., Samelson, L. E. & Sommers, C. L. The LAT story: a tale of cooperativity, coordination, and choreography. Cold Spring Harb. Perspect. Biol. 2, a005512 (2010).
Lo, W. L. et al. Lck promotes Zap70-dependent LAT phosphorylation by bridging Zap70 to LAT. Nat. Immunol. 19, 733–741 (2018).
Wu, J., Motto, D. G., Koretzky, G. A. & Weiss, A. Vav and SLP-76 interact and functionally cooperate in IL-2 gene activation. Immunity 4, 593–602 (1996).
Zhang, W., Sloan-Lancaster, J., Kitchen, J., Trible, R. P. & Samelson, L. E. LAT: the ZAP-70 tyrosine kinase substrate that links T cell receptor to cellular activation. Cell 92, 83–92 (1998).
Bogin, Y., Ainey, C., Beach, D. & Yablonski, D. SLP-76 mediates and maintains activation of the Tec family kinase ITK via the T cell antigen receptor-induced association between SLP-76 and ITK. Proc. Natl Acad. Sci. USA 104, 6638–6643 (2007).
Granja, C., Lin, L. L., Yunis, E. J., Relias, V. & Dasgupta, J. D. PLCγ1, a possible mediator of T cell receptor function. J. Biol. Chem. 266, 16277–16280 (1991).
Bunnell, S. C. et al. Biochemical interactions integrating Itk with the T cell receptor-initiated signaling cascade. J. Biol. Chem. 275, 2219–2230 (2000).
Scharenberg, A. M. & Kinet, J. P. PtdIns-3,4,5-P3: a regulatory nexus between tyrosine kinases and sustained calcium signals. Cell 94, 5–8 (1998).
Clipstone, N. A. & Crabtree, G. R. Identification of calcineurin as a key signalling enzyme in T-lymphocyte activation. Nature 357, 695–697 (1992).
Monks, C. R., Kupfer, H., Tamir, I., Barlow, A. & Kupfer, A. Selective modulation of protein kinase C-Θ during T-cell activation. Nature 385, 83–86 (1997).
Carrasco, S. & Merida, I. Diacylglycerol-dependent binding recruits PKCθ and RasGRP1 C1 domains to specific subcellular localizations in living T lymphocytes. Mol. Biol. Cell 15, 2932–2942 (2004).
Acuto, O. & Michel, F. CD28-mediated co-stimulation: a quantitative support for TCR signalling. Nat. Rev. Immunol. 3, 939–951 (2003).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K. L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat. Cell Biol. 4, 648–657 (2002).
Vander Haar, E., Lee, S. I., Bandhakavi, S., Griffin, T. J. & Kim, D. H. Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat. Cell Biol. 9, 316–323 (2007).
Boomer, J. S. & Green, J. M. An enigmatic tail of CD28 signaling. Cold Spring Harb. Perspect. Biol. 2, a002436 (2010).
Hutloff, A. et al. ICOS is an inducible T-cell co-stimulator structurally and functionally related to CD28. Nature 397, 263–266 (1999).
Yoshinaga, S. K. et al. Characterization of a new human B7-related protein: B7RP-1 is the ligand to the co-stimulatory protein ICOS. Int. Immunol. 12, 1439–1447 (2000).
Rudd, C. E. & Schneider, H. Unifying concepts in CD28, ICOS and CTLA4 co-receptor signalling. Nat. Rev. Immunol. 3, 544–556 (2003).
Zola, H. et al. CD molecules 2005: human cell differentiation molecules. Blood 106, 3123–3126 (2005).
Schober, T. et al. A human immunodeficiency syndrome caused by mutations in CARMIL2. Nat. Commun. 8, 14209 (2017).
Bidere, N. et al. Casein kinase 1α governs antigen-receptor-induced NF-κB activation and human lymphoma cell survival. Nature 458, 92–96 (2009).
Ruland, J. & Hartjes, L. CARD-BCL-10-MALT1 signalling in protective and pathological immunity. Nat. Rev. Immunol. 19, 118–134 (2019).
Delgoffe, G. M. et al. The mTOR kinase differentially regulates effector and regulatory T cell lineage commitment. Immunity 30, 832–844 (2009).
Lin, J. T. et al. Naive CD4 T cell proliferation is controlled by mammalian target of rapamycin regulation of GRAIL expression. J. Immunol. 182, 5919–5928 (2009).
Finlay, D. K. et al. PDK1 regulation of mTOR and hypoxia-inducible factor 1 integrate metabolism and migration of CD8+ T cells. J. Exp. Med. 209, 2441–2453 (2012).
Sun, L., Su, Y., Jiao, A., Wang, X. & Zhang, B. T cells in health and disease. Signal. Transduct. Target. Ther. 8, 235 (2023).
Xue, C. et al. Evolving cognition of the JAK-STAT signaling pathway: autoimmune disorders and cancer. Signal. Transduct. Target. Ther. 8, 204 (2023).
Iida, T. et al. Regulation of cell surface expression of CTLA-4 by secretion of CTLA-4-containing lysosomes upon activation of CD4+ T cells. J. Immunol. 165, 5062–5068 (2000).
Boussiotis, V. A. Molecular and biochemical aspects of the PD-1 checkpoint pathway. N. Engl. J. Med. 375, 1767–1778 (2016).
Lotze, M. T., Olejniczak, S. H. & Skokos, D. CD28 co-stimulation: novel insights and applications in cancer immunotherapy. Nat. Rev. Immunol. 24, 878–895 (2024).
Yokosuka, T. et al. Programmed cell death 1 forms negative costimulatory microclusters that directly inhibit T cell receptor signaling by recruiting phosphatase SHP2. J. Exp. Med. 209, 1201–1217 (2012).
Hui, E. et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 355, 1428–1433 (2017).
Sheppard, K. A. et al. PD-1 inhibits T-cell receptor induced phosphorylation of the ZAP70/CD3ζ signalosome and downstream signaling to PKCθ. FEBS Lett. 574, 37–41 (2004).
Rowshanravan, B., Halliday, N. & Sansom, D. M. CTLA-4: a moving target in immunotherapy. Blood 131, 58–67 (2018).
Collins, A. V. et al. The interaction properties of costimulatory molecules revisited. Immunity 17, 201–210 (2002).
Qureshi, O. S. et al. Trans-endocytosis of CD80 and CD86: a molecular basis for the cell-extrinsic function of CTLA-4. Science 332, 600–603 (2011).
Hedrick, S. M., Cohen, D. I., Nielsen, E. A. & Davis, M. M. Isolation of cDNA clones encoding T cell-specific membrane-associated proteins. Nature 308, 149–153 (1984).
Yanagi, Y. et al. A human T cell-specific cDNA clone encodes a protein having extensive homology to immunoglobulin chains. Nature 308, 145–149 (1984).
Minden, M. D. & Mak, T. W. The structure of the T cell antigen receptor genes in normal and malignant T cells. Blood 68, 327–336 (1986).
Liu, X., Ning, J., Liu, X. & Chan, W. C. J. Mutations affecting genes in the proximal T-cell receptor signaling pathway in peripheral T-cell lymphoma. Cancers 14, 3716 (2022).
Streubel, B., Vinatzer, U., Willheim, M., Raderer, M. & Chott, A. Novel t(5;9)(q33;q22) fuses ITK to SYK in unspecified peripheral T-cell lymphoma. Leukemia 20, 313–318 (2006). This study reported the first gene fusions involving a regulator of TCR signalling in T-NHL.
Attygalle, A. D., Feldman, A. L. & Dogan, A. ITK/SYK translocation in angioimmunoblastic T-cell lymphoma. Am. J. Surg. Pathol. 37, 1456–1457 (2013).
Pechloff, K. et al. The fusion kinase ITK-SYK mimics a T cell receptor signal and drives oncogenesis in conditional mouse models of peripheral T cell lymphoma. J. Exp. Med. 207, 1031–1044 (2010). This study was the first to show that enforced oncogenic signalling within TCR pathways can drive T-NHL formation in vivo.
Dierks, C. et al. The ITK-SYK fusion oncogene induces a T-cell lymphoproliferative disease in mice mimicking human disease. Cancer Res. 70, 6193–6204 (2010).
Debackere, K. et al. Fusion transcripts FYN-TRAF3IP2 and KHDRBS1-LCK hijack T cell receptor signaling in peripheral T-cell lymphoma, not otherwise specified. Nat. Commun. 12, 3705 (2021).
Lemonnier, F. et al. Recurrent TET2 mutations in peripheral T-cell lymphomas correlate with TFH-like features and adverse clinical parameters. Blood 120, 1466–1469 (2012).
Moon, C. S. et al. FYN-TRAF3IP2 induces NF-κB signaling-driven peripheral T cell lymphoma. Nat. Cancer 2, 98–113 (2021). This study determined the mechanism of the FYN–TRAF3IP2 oncogenic fusion in T-NHL and its effect on TCR signalling pathways.
Palomero, T. et al. Recurrent mutations in epigenetic regulators, RHOA and FYN kinase in peripheral T cell lymphomas. Nat. Genet. 46, 166–170 (2014). This study, together with Yoo et al. (2014) and Nagata et al. (2016), identified the recurrent RHOA G17V hotspot mutation in human T-NHL.
Kataoka, K. et al. Integrated molecular analysis of adult T cell leukemia/lymphoma. Nat. Genet. 47, 1304–1315 (2015). This study defined the genomic landscapes of ATLL and reported numerous gain-of-function alterations in TCR and co-receptor pathways.
Tybulewicz, V. L., Ardouin, L., Prisco, A. & Reynolds, L. F. Vav1: a key signal transducer downstream of the TCR. Immunol. Rev. 192, 42–52 (2003).
Chrencik, J. E. et al. Structural basis of guanine nucleotide exchange mediated by the T-cell essential Vav1. J. Mol. Biol. 380, 828–843 (2008).
Fischer, K. D. et al. Vav is a regulator of cytoskeletal reorganization mediated by the T-cell receptor. Curr. Biol. 8, 554–562 (1998).
Morrish, E. et al. The fusion oncogene VAV1-MYO1F triggers aberrant T-cell receptor signaling in vivo and drives peripheral T-cell lymphoma in mice. Eur. J. Immunol. 53, e2250147 (2023). This study, together with Cortes et al. (2022), showed that the VAV1–MYO1F fusion activates TCR signalling and drives T-NHL in vivo.
Abate, F. et al. Activating mutations and translocations in the guanine exchange factor VAV1 in peripheral T-cell lymphomas. Proc. Natl Acad. Sci. USA 114, 764–769 (2017).
Fujisawa, M. et al. Activation of RHOA-VAV1 signaling in angioimmunoblastic T-cell lymphoma. Leukemia 32, 694–702 (2018).
Cortes, J. R. et al. Oncogenic Vav1-Myo1f induces therapeutically targetable macrophage-rich tumor microenvironment in peripheral T cell lymphoma. Cell Rep. 39, 110695 (2022).
Margolis, B. et al. Tyrosine phosphorylation of vav proto-oncogene product containing SH2 domain and transcription factor motifs. Nature 356, 71–74 (1992).
Crespo, P., Schuebel, K. E., Ostrom, A. A., Gutkind, J. S. & Bustelo, X. R. Phosphotyrosine-dependent activation of Rac-1 GDP/GTP exchange by the vav proto-oncogene product. Nature 385, 169–172 (1997).
Han, J. et al. Lck regulates Vav activation of members of the Rho family of GTPases. Mol. Cell Biol. 17, 1346–1353 (1997).
Yoo, H. Y. et al. A recurrent inactivating mutation in RHOA GTPase in angioimmunoblastic T cell lymphoma. Nat. Genet. 46, 371–375 (2014).
Cortes, J. R. et al. RHOA G17V induces T follicular helper cell specification and promotes lymphomagenesis. Cancer Cell 33, 259–273.e7 (2018). This study provided in vivo evidence that RHOA G17V disrupts T cell differentiation and promotes T-NHL development.
Zhang, B. & Zheng, Y. Regulation of RhoA GTP hydrolysis by the GTPase-activating proteins p190, p50RhoGAP, Bcr, and 3BP-1. Biochemistry 37, 5249–5257 (1998).
Zhang, B., Chernoff, J. & Zheng, Y. Interaction of Rac1 with GTPase-activating proteins and putative effectors. A comparison with Cdc42 and RhoA. J. Biol. Chem. 273, 8776–8782 (1998).
Zhang, Y., Cheng, K. & Choi, J. TCR pathway mutations in mature T cell lymphomas. J. Immunol. 211, 1450–1458 (2023).
Robles-Valero, J. et al. Cancer-associated mutations in VAV1 trigger variegated signaling outputs and T-cell lymphomagenesis. EMBO J. 40, e108125 (2021).
Nagata, Y. et al. Variegated RHOA mutations in adult T-cell leukemia/lymphoma. Blood 127, 596–604 (2016).
Park, J. et al. Genomic analysis of 220 CTCLs identifies a novel recurrent gain-of-function alteration in RLTPR (p.Q575E). Blood 130, 1430–1440 (2017).
Leca, J. et al. IDH2 and TET2 mutations synergize to modulate T follicular helper cell functional interaction with the AITL microenvironment. Cancer Cell 41, 323–339.e10 (2023).
Stone, E. L. et al. ICOS coreceptor signaling inactivates the transcription factor FOXO1 to promote Tfh cell differentiation. Immunity 42, 239–251 (2015).
Zang, S. et al. Mutations in 5-methylcytosine oxidase TET2 and RhoA cooperatively disrupt T cell homeostasis. J. Clin. Invest. 127, 2998–3012 (2017).
Patel, V. M. et al. Frequent and persistent PLCG1 mutations in Sézary cells directly enhance PLCγ1 activity and stimulate NFκB, AP-1, and NFAT signaling. J. Invest. Dermatol. 140, 380–389.e4 (2020).
Nicolae, A. et al. Nodal cytotoxic peripheral T-cell lymphoma occurs frequently in the clinical setting of immunodysregulation and is associated with recurrent epigenetic alterations. Mod. Pathol. 35, 1126–1136 (2022).
Gresset, A., Hicks, S. N., Harden, T. K. & Sondek, J. Mechanism of phosphorylation-induced activation of phospholipase C-γ isozymes. J. Biol. Chem. 285, 35836–35847 (2010).
Vaque, J. P. et al. PLCG1 mutations in cutaneous T-cell lymphomas. Blood 123, 2034–2043 (2014).
Wang, L. et al. Genomic profiling of Sezary syndrome identifies alterations of key T cell signaling and differentiation genes. Nat. Genet. 47, 1426–1434 (2015). This study, together with Choi et al. (2015) and da Silva Almeida et al. (2015), characterized the genomic landscapes of CTCL/Sézary syndrome and described multiple alterations in T cell signalling molecules.
Vallois, D. et al. Activating mutations in genes related to TCR signaling in angioimmunoblastic and other follicular helper T-cell-derived lymphomas. Blood 128, 1490–1502 (2016). This study reported the mutational landscapes of TFH cell-derived T-NHL, identifying multiple alterations in TCR and co-receptor signalling molecules.
Coudronniere, N., Villalba, M., Englund, N. & Altman, A. NF-κB activation induced by T cell receptor/CD28 costimulation is mediated by protein kinase C-θ. Proc. Natl Acad. Sci. USA 97, 3394–3399 (2000).
Choi, J. et al. Genomic landscape of cutaneous T cell lymphoma. Nat. Genet. 47, 1011–1019 (2015).
Watatani, Y. et al. Molecular heterogeneity in peripheral T-cell lymphoma, not otherwise specified revealed by comprehensive genetic profiling. Leukemia 33, 2867–2883 (2019).
Garcia-Diaz, N. et al. PLCγ1/PKCθ downstream signaling controls cutaneous T-cell lymphoma development and progression. J. Invest. Dermatol. 142, 1391–1400.e15 (2022).
Kogure, Y. et al. Whole-genome landscape of adult T-cell leukemia/lymphoma. Blood 139, 967–982 (2022).
Woollard, W. J. et al. Candidate driver genes involved in genome maintenance and DNA repair in Sezary syndrome. Blood 127, 3387–3397 (2016).
Baier-Bitterlich, G. et al. Protein kinase C-θ isoenzyme selective stimulation of the transcription factor complex AP-1 in T lymphocytes. Mol. Cell Biol. 16, 1842–1850 (1996).
Sekulic, A. et al. Personalized treatment of Sezary syndrome by targeting a novel CTLA4:CD28 fusion. Mol. Genet. Genomic Med. 3, 130–136 (2015). This study identified gain-of-function CTLA4–CD28 co-receptor fusions and demonstrated their responsiveness to ligand inhibition in a patient.
Rohr, J. et al. Recurrent activating mutations of CD28 in peripheral T-cell lymphomas. Leukemia 30, 1062–1070 (2016).
Yoo, H. Y. et al. Frequent CTLA4-CD28 gene fusion in diverse types of T-cell lymphoma. Haematologica 101, 757–763 (2016).
Egen, J. G. & Allison, J. P. Cytotoxic T lymphocyte antigen-4 accumulation in the immunological synapse is regulated by TCR signal strength. Immunity 16, 23–35 (2002).
Lee, G. J. et al. Mice transgenic for human CTLA4-CD28 fusion gene show proliferation and transformation of ATLL-like and AITL-like T cells. Oncoimmunology 11, 2015170 (2022).
Khayyamian, S. et al. ICOS-ligand, expressed on human endothelial cells, costimulates Th1 and Th2 cytokine secretion by memory CD4+ T cells. Proc. Natl Acad. Sci. USA 99, 6198–6203 (2002).
Vallois, D. et al. RNA fusions involving CD28 are rare in peripheral T-cell lymphomas and concentrate mainly in those derived from follicular helper T cells. Haematologica 103, e360–e363 (2018).
Yoshida, N. et al. Genomic landscape of young ATLL patients identifies frequent targetable CD28 fusions. Blood 135, 1467–1471 (2020).
Marafioti, T. et al. The inducible T-cell co-stimulator molecule is expressed on subsets of T cells and is a new marker of lymphomas of T follicular helper cell-derivation. Haematologica 95, 432–439 (2010).
Amatore, F. et al. ICOS is widely expressed in cutaneous T-cell lymphoma, and its targeting promotes potent killing of malignant cells. Blood Adv. 4, 5203–5214 (2020).
Tafuri, A. et al. ICOS is essential for effective T-helper-cell responses. Nature 409, 105–109 (2001).
Akiba, H. et al. The role of ICOS in the CXCR5+ follicular B helper T cell maintenance in vivo. J. Immunol. 175, 2340–2348 (2005).
Choi, Y. S. et al. ICOS receptor instructs T follicular helper cell versus effector cell differentiation via induction of the transcriptional repressor Bcl6. Immunity 34, 932–946 (2011).
Roncagalli, R. et al. The scaffolding function of the RLTPR protein explains its essential role for CD28 co-stimulation in mouse and human T cells. J. Exp. Med. 213, 2437–2457 (2016).
Uchida, Y. et al. RLTPR Q575E: a novel recurrent gain-of-function mutation in patients with adult T-cell leukemia/lymphoma. Eur. J. Haematol. 106, 221–229 (2021).
da Silva Almeida, A. C. et al. The mutational landscape of cutaneous T cell lymphoma and Sézary syndrome. Nat. Genet. 47, 1465–1470 (2015).
Kameda, T. et al. CARD11 mutation and HBZ expression induce lymphoproliferative disease and adult T-cell leukemia/lymphoma. Commun. Biol. 5, 1309 (2022).
Sommer, K. et al. Phosphorylation of the CARMA1 linker controls NF-κB activation. Immunity 23, 561–574 (2005).
Stinson, J. R. et al. Gain-of-function mutations in CARD11 promote enhanced aggregation and idiosyncratic signalosome assembly. Cell Immunol. 353, 104129 (2020).
Garcia, J. et al. Naturally occurring T cell mutations enhance engineered T cell therapies. Nature 626, 626–634 (2024).
D’Andrea, A. et al. Production of natural killer cell stimulatory factor (interleukin 12) by peripheral blood mononuclear cells. J. Exp. Med. 176, 1387–1398 (1992).
Seder, R. A., Paul, W. E., Davis, M. M. & Fazekas de St Groth, B. The presence of interleukin 4 during in vitro priming determines the lymphokine-producing potential of CD4+ T cells from T cell receptor transgenic mice. J. Exp. Med. 176, 1091–1098 (1992).
Matikainen, S. et al. Interferon-α activates multiple STAT proteins and upregulates proliferation-associated IL-2Rα, c-myc, and pim-1 genes in human T cells. Blood 93, 1980–1991 (1999).
Ross, S. H. & Cantrell, D. A. Signaling and function of interleukin-2 in T lymphocytes. Annu. Rev. Immunol. 36, 411–433 (2018).
Saraiva, M., Vieira, P. & O’Garra, A. Biology and therapeutic potential of interleukin-10. J. Exp. Med. 217, e20190418 (2020).
Fueyo-Gonzalez, F. et al. Interferon-β acts directly on T cells to prolong allograft survival by enhancing regulatory T cell induction through Foxp3 acetylation. Immunity 55, 459–474.e7 (2022).
Lee, K. et al. Primary cytotoxic T-cell lymphomas harbor recurrent targetable alterations in the JAK-STAT pathway. Blood 138, 2435–2440 (2021). This study identified gain-of-function alterations in JAK2–STAT signalling in cytotoxic T cell lymphoma and demonstrated their druggability.
Velusamy, T. et al. A novel recurrent NPM1-TYK2 gene fusion in cutaneous CD30-positive lymphoproliferative disorders. Blood 124, 3768–3771 (2014).
Kuravi, S. et al. Functional characterization of NPM1-TYK2 fusion oncogene. NPJ Precis. Oncol. 6, 3 (2022).
Liang, H. C. et al. The NPM1-TYK2 chimeric fusion promotes activation of STAT family signaling, skewing towards Tfh functional subset differentiation and mature T-cell lymphomagenesis. Blood 140, 9281–9282 (2022).
Crescenzo, R. et al. Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell 27, 516–532 (2015).
Odejide, O. et al. A targeted mutational landscape of angioimmunoblastic T-cell lymphoma. Blood 123, 1293–1296 (2014).
James, C. et al. A unique clonal JAK2 mutation leading to constitutive signalling causes polycythaemia vera. Nature 434, 1144–1148 (2005).
Nishanth, G. et al. Gain of function in Jak2(V617F)-positive T-cells. Leukemia 31, 1000–1003 (2017).
Bahar, B., Barton, K. & Kini, A. R. The role of the exon 13 G571S JAK2 mutation in myeloproliferative neoplasms. Leuk. Res. Rep. 6, 27–28 (2016).
McGirt, L. Y. et al. Whole-genome sequencing reveals oncogenic mutations in mycosis fungoides. Blood 126, 508–519 (2015).
Schatz, J. H. et al. Targeted mutational profiling of peripheral T-cell lymphoma not otherwise specified highlights new mechanisms in a heterogeneous pathogenesis. Leukemia 29, 237–241 (2015).
Bellanger, D. et al. Recurrent JAK1 and JAK3 somatic mutations in T-cell prolymphocytic leukemia. Leukemia 28, 417–419 (2014).
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012).
Koskela, H. L. et al. Somatic STAT3 mutations in large granular lymphocytic leukemia. N. Engl. J. Med. 366, 1905–1913 (2012).
Kiel, M. J. et al. Genomic analyses reveal recurrent mutations in epigenetic modifiers and the JAK-STAT pathway in Sezary syndrome. Nat. Commun. 6, 8470 (2015).
Koo, G. C. et al. Janus kinase 3-activating mutations identified in natural killer/T-cell lymphoma. Cancer Discov. 2, 591–597 (2012).
Waldmann, T. A. & Chen, J. Disorders of the JAK/STAT pathway in T cell lymphoma pathogenesis: implications for immunotherapy. Annu. Rev. Immunol. 35, 533–550 (2017).
Heim, M. H., Kerr, I. M., Stark, G. R. & Darnell, J. E. Jr. Contribution of STAT SH2 groups to specific interferon signaling by the Jak-STAT pathway. Science 267, 1347–1349 (1995).
Kiel, M. J. et al. Integrated genomic sequencing reveals mutational landscape of T-cell prolymphocytic leukemia. Blood 124, 1460–1472 (2014).
Kucuk, C. et al. Activating mutations of STAT5B and STAT3 in lymphomas derived from γδ-T or NK cells. Nat. Commun. 6, 6025 (2015).
Rajala, H. L. et al. Discovery of somatic STAT5b mutations in large granular lymphocytic leukemia. Blood 121, 4541–4550 (2013).
Grabstein, K. H. et al. Cloning of a T cell growth factor that interacts with the β chain of the interleukin-2 receptor. Science 264, 965–968 (1994).
Tan, J. T. et al. IL-7 is critical for homeostatic proliferation and survival of naive T cells. Proc. Natl Acad. Sci. USA 98, 8732–8737 (2001).
Schrader, A. et al. Actionable perturbations of damage responses by TCL1/ATM and epigenetic lesions form the basis of T-PLL. Nat. Commun. 9, 697 (2018).
Herling, M. et al. High TCL1 expression and intact T-cell receptor signaling define a hyperproliferative subset of T-cell prolymphocytic leukemia. Blood 111, 328–337 (2008).
Chen, Y. W. et al. Receptor-type tyrosine-protein phosphatase κ directly targets STAT3 activation for tumor suppression in nasal NK/T-cell lymphoma. Blood 125, 1589–1600 (2015).
Kleppe, M. et al. Mutation analysis of the tyrosine phosphatase PTPN2 in Hodgkin’s lymphoma and T-cell non-Hodgkin’s lymphoma. Haematologica 96, 1723–1727 (2011).
Simoncic, P. D., Lee-Loy, A., Barber, D. L., Tremblay, M. L. & McGlade, C. J. The T cell protein tyrosine phosphatase is a negative regulator of Janus family kinases 1 and 3. Curr. Biol. 12, 446–453 (2002).
You-Ten, K. E. et al. Impaired bone marrow microenvironment and immune function in T cell protein tyrosine phosphatase-deficient mice. J. Exp. Med. 186, 683–693 (1997).
Wiede, F. et al. T cell protein tyrosine phosphatase attenuates T cell signaling to maintain tolerance in mice. J. Clin. Invest. 121, 4758–4774 (2011).
Karaca Atabay, E. et al. Tyrosine phosphatases regulate resistance to ALK inhibitors in ALK+ anaplastic large cell lymphoma. Blood 139, 717–731 (2022).
Wartewig, T. & Ruland, J. PD-1 tumor suppressor signaling in T cell lymphomas. Trends Immunol. 40, 403–414 (2019).
Wartewig, T. et al. PD-1 is a haploinsufficient suppressor of T cell lymphomagenesis. Nature 552, 121–125 (2017). This study identified PD1 as a tumour suppressor in T-NHL and showed that it is frequently mutated in human cases.
Park, J. et al. Integrated genomic analyses of cutaneous T-cell lymphomas reveal the molecular bases for disease heterogeneity. Blood 138, 1225–1236 (2021). This study demonstrated that loss of PD1 has prognostic significance in human cutaneous T-NHL.
Wartewig, T. et al. PD-1 instructs a tumor-suppressive metabolic program that restricts glycolysis and restrains AP-1 activity in T cell lymphoma. Nat. Cancer 4, 1508–1525 (2023). This study identified PD1-dependent tumour-suppressor regulation of metabolic and epigenetic programs in T-NHL.
Ratner, L., Waldmann, T. A., Janakiram, M. & Brammer, J. E. Rapid progression of adult T-cell leukemia-lymphoma after PD-1 inhibitor therapy. N. Engl. J. Med. 378, 1947–1948 (2018). This study was the first to report hyperprogression of human T-NHL after pharmacological PD1 checkpoint.
Rauch, D. A. et al. Rapid progression of adult T-cell leukemia/lymphoma as tumor-infiltrating Tregs after PD-1 blockade. Blood 134, 1406–1414 (2019).
Gao, Y. et al. Hyperprogression of cutaneous T cell lymphoma after anti-PD-1 treatment. JCI Insight 8, e164793 (2023).
Anand, K. et al. T-cell lymphoma secondary to checkpoint inhibitor therapy. J Immunother. Cancer 8, e000104 (2020).
Ungewickell, A. et al. Genomic analysis of mycosis fungoides and Sezary syndrome identifies recurrent alterations in TNFR2. Nat. Genet. 47, 1056–1060 (2015).
Wertz, I. E. et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-κB signalling. Nature 430, 694–699 (2004).
Coornaert, B. et al. T cell antigen receptor stimulation induces MALT1 paracaspase-mediated cleavage of the NF-κB inhibitor A20. Nat. Immunol. 9, 263–271 (2008).
Takagi, M. et al. Haploinsufficiency of TNFAIP3 (A20) by germline mutation is involved in autoimmune lymphoproliferative syndrome. J. Allergy Clin. Immunol. 139, 1914–1922 (2017).
Braun, F. C. et al. Tumor suppressor TNFAIP3 (A20) is frequently deleted in Sezary syndrome. Leukemia 25, 1494–1501 (2011).
Ahn, H., Yang, J. M., Jeon, Y. K. & Paik, J. H. Clinicopathologic implications of TNFAIP3/A20 deletions in extranodal NK/T-cell lymphoma. Genes Chromosomes Cancer 57, 231–239 (2018).
Chen, C. et al. TNFAIP3 mutation may be associated with favorable overall survival for patients with T-cell lymphoma. Cancer Cell Int. 21, 490 (2021).
Wang, C. et al. IDH2R172 mutations define a unique subgroup of patients with angioimmunoblastic T-cell lymphoma. Blood 126, 1741–1752 (2015).
Mao, X. et al. Amplification and overexpression of JUNB is associated with primary cutaneous T-cell lymphomas. Blood 101, 1513–1519 (2003).
Byrd, J. C. et al. A phase I, first-in-human study of ONO-7018 in patients with relapsed/refractory non-Hodgkin lymphoma or chronic lymphocytic leukemia. J. Clin. Oncol. https://doi.org/10.1200/JCO.2024.42.16_suppl.TPS708 (2024).
Nguyen, T. B. et al. Dasatinib is an effective treatment for angioimmunoblastic T-cell lymphoma. Cancer Res. 80, 1875–1884 (2020).
Kumar, A. et al. Pilot trial of ibrutinib in patients with relapsed or refractory T-cell lymphoma. Blood Adv. 2, 871–876 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/study/NCT03372057 (2025).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/study/NCT03770000 (2022).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/study/NCT03952078 (2019).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/study/NCT02567656 (2015).
Iyer, S. P. et al. Final results of phase 1/1b study of tenalisib, dual PI3K δ/γ inhibitor in patients with relapsed/refractory T-cell lymphoma. Blood 134, 2831 (2019).
Braun, T. et al. Multiomic profiling of T cell lymphoma after therapy with anti-BCMA CAR T cells and GPRC5D-directed bispecific antibody. Nat. Med. 31, 1145–1153 (2025).
Perica, K. et al. CD4+ T-cell lymphoma harboring a chimeric antigen receptor integration in TP53. N. Engl. J. Med. 392, 577–583 (2025).
Jain, N. et al. TET2 guards against unchecked BATF3-induced CAR T cell expansion. Nature 615, 315–322 (2023).
Manso, R. et al. Recurrent presence of the PLCG1 S345F mutation in nodal peripheral T-cell lymphomas. Haematologica 100, e25–e27 (2015).
Hornakova, T. et al. Acute lymphoblastic leukemia-associated JAK1 mutants activate the Janus kinase/STAT pathway via interleukin-9 receptor α homodimers. J. Biol. Chem. 284, 6773–6781 (2009).
Malinge, S. et al. Activating mutations in human acute megakaryoblastic leukemia. Blood 112, 4220–4226 (2008).
Lobello, C. et al. STAT3 and TP53 mutations associate with poor prognosis in anaplastic large cell lymphoma. Leukemia 35, 1500–1505 (2021).