Drug Preparation
VLPs (Bendao Gene Co., Ltd., China) and lentiviral empty vectors (noload) were dissolved in phosphate-buffered saline (PBS, pH 7.2) and subsequently sterilized by filtration using a 0.22 μm filter prior to use.
Animal grouping
A total of 12 male C57BL/6 wild-type (WT) mice and 12 male FcµR-/- mice were used in the study, with each group being housed separately. After a one-week acclimatization period, the mice were randomly assigned into six groups (n = 4 per group) as follows: WT mice received injections of either PBS (WT), lentiviral empty vector (WT+noload), or VLPs (WT + VLP); FcµR-/- mice, representing the MZB-deficient group, received PBS (FcµR-/-), lentiviral empty vector (FcµR-/-+noload), or VLPs (FcµR-/-+VLP).
Establishment of a VLP-based immunization model
The first immunization and the second immunization were carried out on day 0 and day 21 respectively. Mice in the six groups received footpad injections of either VLPs, lentiviral empty vector solution (0.1 mg/kg), or an equivalent volume of PBS, according to their group assignment. On day 35, all mice were euthanized via cervical dislocation following anesthesia (1% sodium pentobarbital, 50 mg/kg, intraperitoneal injection) in accordance with the humane standards of the American Veterinary Medical Association (AVMA). Plasma samples were collected via cardiac puncture, and spleen tissues were harvested immediately after euthanasia for subsequent experimental analyses.
Body weight of each mouse was measured using an electronic balance (accuracy: 0.01 g) at two key time points: prior to acclimatization (baseline, day 7) and immediately before euthanasia (day 35). The mean body weight ± standard deviation (SD) of each group is summarized in Table 1, ensuring transparency of animal physiological status throughout the experimental period. No significant differences in baseline body weight were observed among the six groups (P > 0.05, one-way ANOVA), confirming the homogeneity of group allocation.
Table 1 Mean body weight ± SD of mice in each experimental group at baseline and day 35.Detection of mouse SARS-CoV-2 Spike antibody (IgG)
The detection of mouse SARS-CoV-2 Spike-specific IgG antibodies was performed following the manufacturer’s instructions provided with the enzyme-linked immunosorbent assay (ELISA) kit (Vazyme Co., Ltd., China).
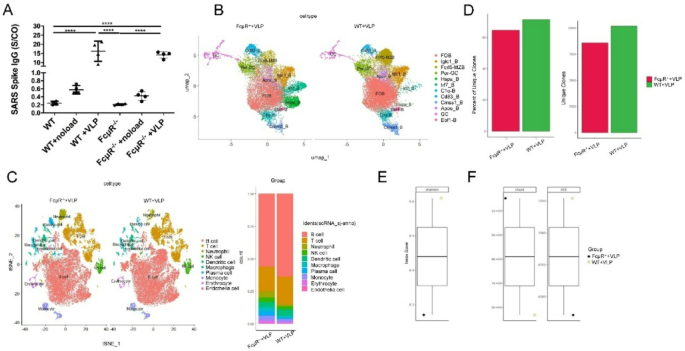
Single-Cell sequencing of mouse spleen cells
Mouse spleens were isolated and single-cell sequencing along with subsequent analysis was conducted by Shanghai Labway Clinical Laboratory Co., Ltd.
BCR repertoire analysis
The BCR repertoire analysis included four core indicators: unique BCR clone counts, Shannon entropy, Chao1 index, and ACE index. This analysis was performed by Shanghai Labway Clinical Laboratory Co., Ltd. using standardized bioinformatics workflows to ensure data consistency and reliability.
Collection of human experiment samples
2 mL of venous blood were collected from each volunteer into EDTA-anticoagulated tubes and centrifuged at 1500 × g for 10 min. The plasma was collected, heat-inactivated at 56 °C for 30 min, aliquoted, and stored at − 80 °C. The remaining cellular fraction was resuspended in PBS and PBMCs were isolated from red blood cells using Ficoll-Paque™ Plus (Cytiva, USA) in accordance with the manufacturer’s instructions.
Detection of SARS-CoV-2, IgG and IgM antibodies
SARS-CoV-2 IgG and total antibody levels in plasma were measured using the Caris 2000 fully automated chemiluminescence analyzer (Xiamen Wantai Co.) along with corresponding reagents based on chemiluminescent microparticle immunoassay. Samples with a signal-to-cutoff (S/CO) ratio ≥ 1.0 COI were considered positive.
Detection of blood routine indicators
Routine blood parameters were measured using the BC-6800Plus automated hematology analyzer (Shenzhen Mindray Co.) along with the corresponding reagents. The assessed indicators included white blood cell (WBC) count, lymphocyte count, C-reactive protein (CRP), monocyte ratio, eosinophil ratio, basophil ratio, absolute neutrophil count, absolute monocyte count, absolute eosinophil count, absolute basophil count, red blood cell count, hemoglobin level, and platelet count.
Rapid detection of surrogate neutralization activity via RBD–ACE2 competitive binding
The PocRoc® SARS-CoV-2 Neutralizing Antibody Rapid Detection Kit (Lumigenex, China), a RBD–ACE2 competitive binding rapid kit was used to detect the surrogate neutralization activity of plasma antibodies. This assay reflects the ability of antibodies to block the RBD-ACE2 interaction. Each test card was placed on a clean, dry surface and labeled according to the corresponding sample. Thereafter, using a dropper, approximately 40 µL of plasma was added to the sample well, followed by one drop of sample buffer. A timer was started, and results were read precisely 15 min later. This assay operates on a competitive binding principle, with the results expressed as the RBD test/control ratio, which is inversely correlated with the surrogate neutralization titer.
Microarray- based plasma peptide chip analysis
A 7-chamber or 14-chamber rubber gasket was mounted onto each slide to create separate compartments for identical sub-arrays. As previously described, these microarray chips were used for plasma analysis. Slides stored at − 80 °C were brought to room temperature, then incubated in blocking buffer (3% BSA in 1× PBS containing 0.1% Tween-20) for 3 h. Each sub-array was then incubated with diluted plasma or antibody: 400 µL for 7-sub-array chambers and 200 µL for 14-sub-array chambers, for 2 h. For most samples, plasma was diluted at 1:200, and free peptides were added at a concentration of 0.25 mg/mL. For antibody-enriched samples, 0.1–0.5 µg of antibody was added to 200 µL of incubation buffer. After incubation, the arrays were washed with 1× PBST and incubated for 1 h at room temperature with Cy3-conjugated goat anti-human IgG and Alexa Fluor 647-conjugated donkey anti-human IgM (Jackson ImmunoResearch, PA, USA), each diluted 1:1000 in 1× PBS. Afterward, the slides were washed with 1× PBS, centrifuged, and air-dried at room temperature. The microarrays were scanned using a LuxScan 10 K-A scanner (CapitalBio Corporation, Beijing, China) with settings of 95% laser power/PMT 550 for IgM and 95% laser power/PMT 480 for IgG. Fluorescence intensity was quantified using GenePix Pro 6.0 software (Molecular Devices, CA, USA)18.
Cytokine analysis
Plasma cytokine levels, including IL-1β, IL-2, IL-4, IL-5, IL-6, IL-8, IL-10, IL-12, IL-17, TNF-α, IFN-α, and IFN-γ, were measured using a human cytokine detection kit (RaiseCare Biological Technology Co., Ltd., Qingdao, Shandong, China) based on a flow-based microsphere array. All procedures were performed in accordance with the manufacturer’s instructions. Samples were analyzed using a FACS Canto II flow cytometer (BD Biosciences, USA).
Labeling of Membrane – Surface molecules detected by flow cytometry
Conventional lymphocyte subsets were analyzed by multi-color flow cytometry using the following human monoclonal antibodies (all from BD Biosciences, USA): CD3-FITC, CD4-PE-Cy7, CD8-APC-Cy7, CD19-APC, CD45-PerCP-Cy5.5, CD16-PE and CD56-PE. Surface marker expression was used to determine the proportions and absolute numbers of total T cells (CD45+CD3+), helper T cells (Th, CD3+CD4+), cytotoxic T cells (Tc, CD3+CD8+), natural killer (NK) cells (CD3-CD16+CD56+), and B cells (CD3-CD19+). All samples were processed and analyzed using a BD FACS Canto II flow cytometer, and data were analyzed with FlowJo software (version 10.0). Peripheral blood was collected, and red blood cells were lysed using red blood cell lysis buffer (Sigma-Aldrich, USA). Single-cell suspensions were then prepared for surface staining. Fc receptors were blocked using anti-human CD16/CD32 (BioLegend, CA, USA), followed by staining with a panel of antibodies including: CD19-FITC, CD21-PE-Cy7, CD23-BV510, Live/Dead viability dye -BV421, CD45-APC, CD3-FITC, CD4-BV510, CD8-APC-Cy7, CXCR5-BV421 and ICOS-PE (all from BD Biosciences, USA). Cells were incubated at 4 °C for 30 minutes, washed, resuspended, and analyzed using the BD FACS Canto II flow cytometer. Gating was performed based on isotype control staining profiles.
Statistical methods
Statistical analyses were performed using GraphPad Prism 8 software (GraphPad Software, USA). A p-value of *P < 0.05 was considered statistically significant, while **P < 0.01, ***P < 0.001 and ****P < 0.0001 indicated increasingly higher levels of statistical significance. All tests were two-tailed and data distribution was assessed for normality prior to test selection. In figures “n.s.” indicates non-significance (P ≥ 0.05) and “n.d.” stands for not detected.
In Mouse cohort, six groups with n = 4 per group. Relevant indicators were compared across groups using one-way ANOVA.
In Human vaccine cohort, four independently recruited groups Pre-vaccine (n = 18) Post-2nd dose (7 days) (n = 18 paired with Pre-vaccine) 6 months post-2nd dose (n = 30) and Post-booster (7 days) (n = 30). Paired comparisons between Pre-vaccine and Post-2nd dose were analyzed using the Wilcoxon matched-pairs signed-rank test. Comparisons among three independent post-vaccination groups were performed with the Kruskal–Wallis test followed by Dunn’s post-hoc test with Bonferroni correction.
In Elderly Omicron infection cohort, two groups Acute phase (n = 25) and Recovery phase (n = 38). Hematological parameters antibody levels and immune cell subsets were compared using the Mann–Whitney U test.





![[MBN] A Spring Herb Table to Restore Weakened Immunity A special remedy will be unveiled to tackle chronic fatigue and weakened immunity, the unwelcome guests of the changing seasons. On the program, Venerable Jeokmun, designated as Master No. 3 of Korean Buddhist temple cuisine, faces off in a spring herb cooking contest against "announcer-chef" Cho Woo-jong, who holds a licensed Western cuisine chef qualification. From spring cabbage bibimbap that has been selling out on social networking service (SNS) platforms to seasonal menus infused with the master’s secret techniques, the show presents a spring dining table that satisfies both the palate and health.](https://www.vitaminrush.com/wp-content/uploads/2026/03/20260331_01110129000004_L00-770x515.jpg)