SA induces PsAvh109 expression to synchronize effector deployment
Numerous effector genes of P. sojae are expressed during the early stages of infection, coinciding with the critical window of SA-mediated resistance to biotrophic pathogens. To investigate whether SA functions as an inductive signal for RxLR effector gene expression, P. sojae germ tubes were treated with 100 μM SA. Subsequent RNA sequencing (RNA-seq) revealed that 2608 genes were significantly upregulated and 2993 were downregulated in response to SA treatment compared with ethanol (EtOH) controls (Supplementary Fig. 1a). Among the previously annotated RxLR effectors, 64 exhibited significant upregulated, whereas only six were downregulated (Supplementary Fig. 1b). PsAvh109 was among the most strongly induced effectors and was selected for further investigation due to its demonstrated ability to suppress immune responses triggered by pathogen-associated molecular patterns (PAMPs) such as INF1 and XEG133,34.
To monitor protein accumulation, we generated an in situ complementation strain (C-PsAvh109-6His-mCherry) by replacing the endogenous PsAvh109 gene. Western blot analysis revealed that exogenous SA treatment significantly increased PsAvh109-6His-mCherry protein levels relative to EtOH controls (Supplementary Fig. 1c). Moreover, a comparison of the inductive effects of various exogenous phytohormones, such as SA, JA, and abscisic acid (ABA) demonstrated that SA exerted robust induction of PsAvh109 (Fig. 1a). Furthermore, RT-qPCR analysis confirmed that SA induced PsAvh109 transcription in a concentration-dependent manner (Fig. 1b).
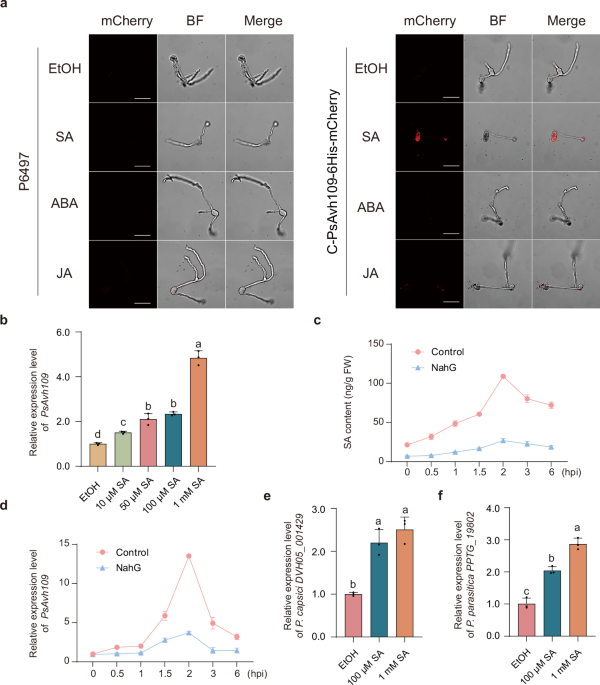
Fig. 1: Salicylic acid induces the production of effector Avh109 in Phytophthora.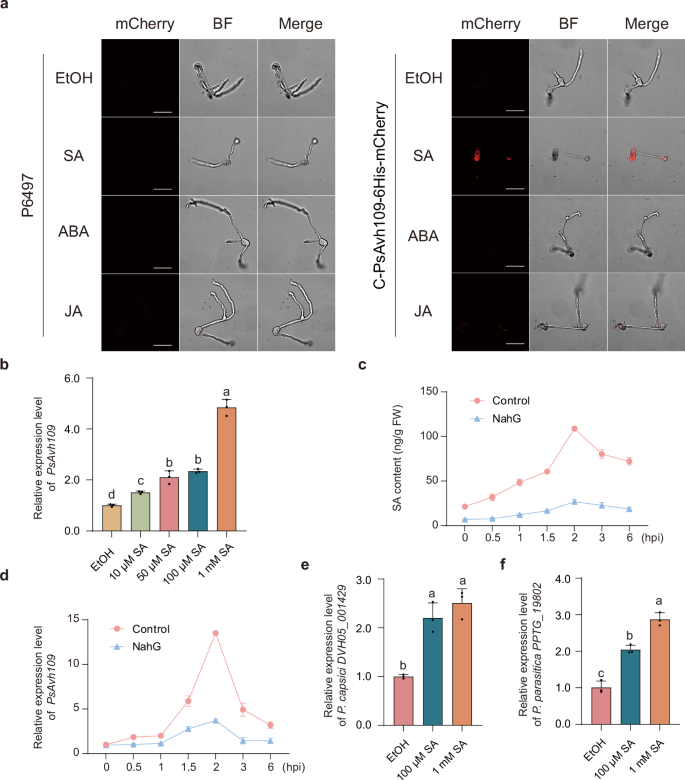 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a Confocal microscopy analysis of Avh109 effector fluorescence in hyphae of P. sojae wild-type strain P6497 and the C-PsAvh109-6His-mCherry gene-edited strain. Hyphae were treated with 100 μM SA, JA, or ABA for 30 min, with ethanol (EtOH) used as a solvent control. Scale bar: 20 μm. b PsAvh109 transcript levels were quantified by RT-qPCR following treatment with SA (10 μM, 50 μM, 100 μM, 1 mM) for 30 min. EtOH served as a solvent control. Data were normalized to the reference gene PsACT and expressed as means ± SD (n = 3). c SA content in root hairs of NahG-overexpressing or control infected with P. sojae. Data are presented as mean ± SD (n = 3). FW, fresh weight. d Transcriptional levels of the P. sojae effector Avh109 in the samples described in (c). Data are presented as mean ± SD (n = 3). e The transcription levels of the DVH05_001429 gene in P. capsici were measured by RT-qPCR following induction with SA for 30 min. EtOH was used as a control. Relative transcription levels were normalized to the reference gene PcUBC. Data are presented as the mean ± SD (n = 3). f The transcription levels of the PPTG_19802 gene in P. parasitica were measured by RT-qPCR following induction with SA for 30 min. EtOH was used as a control. Relative transcription levels were normalized to the reference gene PpACT. Data are presented as the mean ± SD (n = 3). (b,e,f) Different letters indicate significant differences (P < 0.01; one-way ANOVA with Tukey’s multiple comparisons test).
To evaluate the physiological significance of host-derived SA, we utilized transgenic soybean roots expressing the salicylate hydroxylase NahG, which depletes endogenous SA35, alongside control roots. Quantitative analysis via LC-MS/MS revealed a substantial elevation of endogenous SA in infected control roots, rising from approximately 30 ng/g fresh weight (FW) in mock-inoculated tissues to 130 ng/g FW at 2 h post-inoculation (hpi). This transient peak in host SA accumulation was temporally coupled with the maximal transcriptional induction of PsAvh109 (Fig. 1c, d). In NahG-transgenic roots, both SA accumulation and PsAvh109 induction were significantly attenuated relative to controls, establishing a robust positive correlation between host SA titers and effector expression (Fig. 1c, d). Compared to the control, P. sojae infected NahG-transgenic roots exhibited a 1.4-fold increase in oospore production and a 1.8-fold increase in pathogen biomass (Supplementary Fig. 1d–f), underscoring the indispensable role of SA in plant defense against pathogen. Furthermore, homologous of PsAvh109 in P. capsici (DVH05_001429) and P. parasitica (PPTG_19802) also exhibited SA-inducible expression (Fig. 1e, f), suggesting that SA responsiveness is a conserved trait among these oomycete pathogens.
PsAvh109 requires nuclear localization to suppress immunity and promote the virulence of P. sojae
PsAvh109 is highly conserved across various Phytophthora species, including P. infestans, P. parasitica, P. cactorum and P. capsici. Notably, all identified PsAvh109 homologs harbor a predicted nuclear localization signal (NLS) (Fig. 2a and S2). The mature form of PsAvh109 fused with green fluorescent protein (GFP) accumulated in the nuclei of Nicotiana benthamiana epidermal cells, co-localizing with the nuclear marker histone 2B fused to mCherry (H2B-mCherry) (Fig. 2b). To disrupt nuclear localization, we generated NLS mutants (PsAvh109AAA and PsAvh109NES) by either substituting all conserved lysine (K) and arginine (R) residues with alanine (A) or by appending a nuclear export signal (NES) (Fig. 2a). These modifications abolished nuclear targeting, as evidenced by their exclusion from H2B-mCherry labeled nuclei (Fig. 2b). Immunoblotting of nuclear and total protein fractions from N. benthamiana cells confirmed the specific enrichment of PsAvh109-GFP within the nuclear fraction, whereas PsAvh109AAA-GFP and free GFP remained absent (Supplementary Fig. 3a).
Fig. 2: PsAvh109 is a nucleus-localized effector that is essential for P. sojae virulence.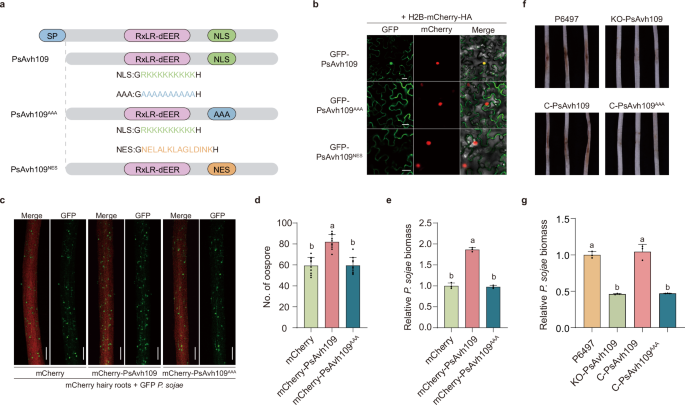 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a Schematic representation of the PsAvh109 protein domain architecture, highlighting the nuclear localization signal (NLS) in green. The PsAvh109AAA mutant contains substitutions of lysine (K) and arginine (R) residues with alanine (A) within the NLS region (highlighted in blue), whereas the PsAvh109NES mutant features a replacement of the NLS with a nuclear export signal (NES), indicated in orange. b Confocal microscopy analysis revealed the subcellular localization of PsAvh109 and its mutants. GFP-PsAvh109 colocalized with the nuclear marker H2B-mCherry after 36 h of co-expression. Scale bar: 20 μm. c Co-inoculation of soybean root hairs expressing mCherry, mCherry-PsAvh109, or mCherry-PsAvh109AAA with a GFP-tagged P. sojae transformant (strain P6497). Fluorescence microscopy images were acquired at 48 h post-inoculation (hpi). Scale bar: 200 μm. d Quantification of P. sojae oospore formation in soybean root hairs. Data are presented as mean ± standard deviation (SD) (n = 10). e Relative P. sojae biomass in soybean root hairs, determined by quantitative PCR (qPCR). Data are presented as mean ± SD (n = 3). f Phenotypic analysis of P. sojae virulence in soybean hypocotyls. Wild-type strain P6497, the PsAvh109 knockout mutant (KO-PsAvh109), and two complemented strains (C-PsAvh109 and C-PsAvh109AAA) were inoculated onto soybean hypocotyls. Representative images were taken at 48 hpi. The experiment was independently repeated three times with similar results. g Quantification of P. sojae biomass in infected soybean hypocotyls by qPCR, based on the ratio of P. sojae to soybean genomic DNA. Data are shown as mean ± SD (n = 3). (d,e,g) Different letters indicate significant differences (P < 0.01; one-way ANOVA with Tukey’s multiple comparisons test).
To directly monitor effector translocation during the infection process, a functional PsAvh109-GFP fusion was expressed in P. sojae. Confocal microscopy revealed prominent GFP signals within the nuclei of soybean cells in immediate proximity to invading hyphae; notably, no such nuclear signal was observed in plants infected with a control strain expressing GFP alone (Supplementary Fig. 3b). Consistently, PsAvh109-GFP was detected in nuclear fractions isolated from infected soybean tissues, providing definitive evidence that PsAvh109 is translocated into the host nucleus during natural infection (Supplementary Fig. 3c).
In soybean hairy roots, the expression of PsAvh109, but not the NLS mutant PsAvh109AAA, significantly enhanced P. sojae virulence, as evidenced by 1.4-fold increase in oospore production and a 1.8-fold increase in pathogen biomass relative to the mCherry control (Fig. 2c–e). Similarly, PsAvh109 promoted P. capsici infection in N. benthamiana, resulting in a 1.3-fold expansion in lesion diameter and a 2.3-fold increase in pathogen biomass (Supplementary Fig. 4a–c). In contrast, the PsAvh109AAA mutant exhibited no such stimulatory effect. Furthermore, PsAvh109 also suppressed INF1-induced programmed cell death (PCD), a suppressive activity that was abolished in the PsAvh109AAA mutant (Supplementary Fig. 4a).
CRISPR/Cas9-mediated knockout of PsAvh109 (KO-PsAvh109) in P. sojae did not impair mycelial growth or zoospore production (Supplementary Fig. 4d–f). However, it markedly attenuated virulence, resulting in reduced lesions on soybean hypocotyls and a 50% decrease in pathogen biomass compared with the wild-type strain. Pathogenicity was fully restored in the complementation strain (C-PsAvh109), whereas the nuclear localization-deficient mutant (C-PsAvh109AAA) failed to rescue the virulence defect (Fig. 2f, g). These data indicate that PsAvh109 is essential for full virulence and that its function is strictly dependent on nuclear localization.
PsAvh109 targets the Mediator subunit MED21 to disrupt defense transcription
Yeast two-hybrid (Y2H) screening identified Glycine max MED21 (GmMED21), a conserved subunit within the middle module of the transcriptional Mediator complex36,37, as a candidate interactor of PsAvh109 (Supplementary Fig. 5). Further analysis revealed that PsAvh109 also associates with the N. benthamiana homolog, NbMED21, indicating a conserved interaction across different host species (Supplementary Fig. 5). Subsequent directed Y2H assays confirmed the specificity of this interaction, demonstrating that PsAvh109 binds to GmMED21 but not to adjacent Mediator subunits (GmMED6, GmMED7, GmMED10, GmMED19, or GmMED25) (Supplementary Fig. 5).
Luciferase complementation imaging (LCI) assays revealed a robust luminescence signal in N. benthamiana leaves co-expressing GmMED21-nLUC and cLUC-PsAvh109; however, this signal was abolished in NLS mutants, confirming the in vivo interaction between GmMED21 and PsAvh109 (Fig. 3a). Co-immunoprecipitation (Co-IP) experiments further corroborated this interaction, as GFP-PsAvh109 effectively co-precipitated GmMED21-mCherry-3HA from N. benthamiana extracts. Consistent with the negative controls, neither GFP-PsAvh109NES nor GFP-PsAvh109AAA was detected in the immunoprecipitated fractions (Fig. 3b). Conversely, in vitro pull-down assays showed that GmMED21-His interacted with GST-PsAvh109, as well as with the GST-PsAvh109NES and GST-PsAvh109AAA mutants (Fig. 3c). These results indicate that PsAvh109 physically interacts with GmMED21 both in vivo and in vitro. Notably, the loss of interaction in vivo for the NLS mutations likely due to altered protein translocation rather than a disruption of the physical binding affinity for GmMED21.
Fig. 3: PsAvh109 targets GmMED21 to modulate host transcription and immunity.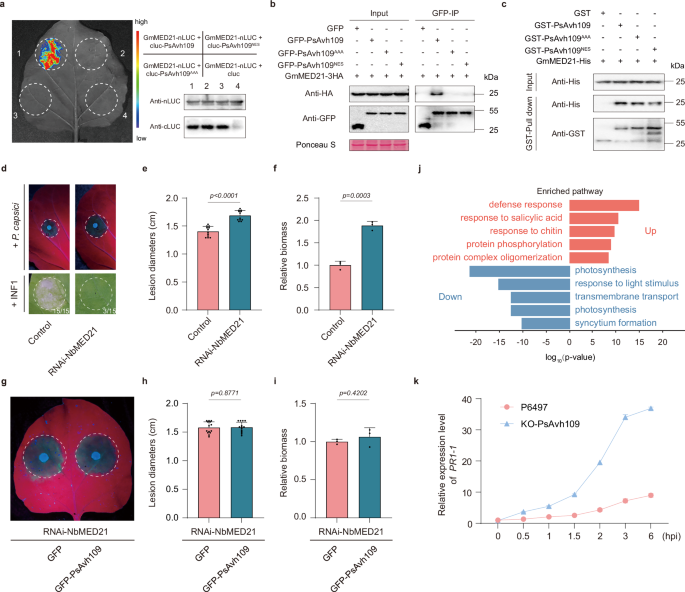 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a LCI assay in N. benthamiana confirming PsAvh109-GmMED21 interaction. Luminescence was detected 2 days post-agrobacterium co-infiltration following D-luciferin application; protein expression was verified by Western blotting. b Co-IP in N. benthamiana validating the PsAvh109-GmMED21 interaction. GFP-tagged PsAvh109 variants were co-expressed with GmMED21-3HA; anti-GFP beads pulled down complexes analyzed by anti-GFP and anti-HA immunoblotting. Ponceau S staining served as loading control; molecular weights (kDa) indicated. c Pull-down assay demonstrated direct binding between GST-PsAvh109 variants immobilized on glutathione resin and His-GmMED21. Input samples and eluates were analyzed using anti-His and anti-GST antibodies. d Silencing of NbMED21 enhances susceptibility to P. capsici and suppresses INF1-triggered cell death compared to RNAi-GFP controls. UV fluorescence shows lesion expansion at 48 hpi (white circles indicate lesion margins); expression of INF1-induced cell death at 36 hpi (numbers indicate the number of leaves exhibiting cell necrosis). e Lesion diameters in (d) quantified as median ± SD (n = 12). f Relative P. capsici biomass in (d) quantified by qPCR. (mean ± SD, n = 3). g Complementation assay: RNAi-NbMED21 leaves expressing GFP or GFP-PsAvh109 were challenged with P. capsici. UV fluorescence shows lesion expansion at 48 hpi. h Lesion diameter in (g) quantified as median ± SD (n = 14). i Relative P. capsici biomass in (g) quantified by qPCR (mean ± SD, n = 3). j Gene Ontology enrichment analysis of PsAvh109-regulated genes. Top enriched biological process terms are shown for up-regulated (red) and down-regulated (blue) gene sets, ranked by fold enrichment. Statistical significance was determined using a two-sided Fisher’s exact test. P-values were adjusted for multiple comparisons via the Benjamini-Hochberg method to control false discovery rate (FDR); q-values are provided in Source Data. The x-axis represents log10 (P-value). k Transcriptional levels of the plant PR1-1 gene in soybean plants infected with the wild-type P. sojae strain P6497 or the KO-PsAvh109 mutant. Expression levels were normalized to GmCYP2 as an internal reference. Data are presented as mean ± SD (n = 3). (e, f, h, i) P values were derived using two-tailed Student’s t-test. (a, b, c) All the experiments were repeated three times with similar results.
To evaluate the roles of MED21 as the effector target of PsAvh109, we performed Agrobacterium-mediated transient expression of INF1 or inoculated P. capsici onto RNAi-NbMED21 N. benthamiana leaves. INF1-induced PCD was significantly attenuated in NbMED21-knockdown tissues (Fig. 3d). Compared with the wild-type (WT) control, silencing of NbMED21 resulted in a 1.2-fold increase in lesion diameter and a two-fold increase in P. capsici biomass in infected leaves (Fig. 3d–f). Notably, the silencing of NbMED21 abolished the virulence-promoting activity of PsAvh109; specifically, as PsAvh109 expression failed to further enhance P. capsici infection in NbMED21-silenced tissues (Fig. 3g). In these silenced tissues, lesion diameter and pathogen biomass were comparable to those observed in the GFP control tissues (Fig. 3g–i). These results suggest that the interaction with MED21 is essential for PsAvh109 to facilitate pathogen infection.
To characterize the transcriptomic changes orchestrated by PsAvh109 during infection, we performed RNA-seq analysis on soybean plants inoculated with either wild-type P. sojae P6497 or an isogenic PsAvh109 knockout mutant (KO-PsAvh109). Comparative transcriptomic profiling revealed a profound shift in the expression landscape of KO-PsAvh109 infected soybean, with 3,518 genes significantly upregulated and 4,562 genes downregulated compared with P6497-infected controls (Supplementary Fig. 6a). Gene Ontology (GO) enrichment analysis of these differentially expressed genes (DEGs) uncovered a significant overrepresentation of defense-related biological processes, notably defense responses, response to SA, and response to chitin (Fig. 3j). Notably, the presence of PsAvh109 led to the potent repression of SA-responsive genes, including seven PR1 homologs (GmPR1-1/2/3/4/5/6/7) that are vital for SA-mediated immunity (Supplementary Fig. 6b). Furthermore, we quantified PR1-1 expression levels in soybean inoculated with either the wild-type strain P6497 or the KO-PsAvh109 mutant. In the early stages of infection, PR1-1 transcription is sharply upregulated in plants inoculated with the KO-PsAvh109 mutant. Conversely, PR1-1 transcript accumulation follows a more gradual trajectory in plants infected with the wild-type strain P6497 (Fig. 3k). Collectively, these findings demonstrate that PsAvh109 suppresses SA-dependent defense transcription, specifically PR1 expression, during compatible host interactions, a process likely driven by its interference with GmMED21.
GmMED21 and GmMED6 act as the molecular switch of SA controlled gene transcription and blocked by PsAvh109 via competitive binding
The head and middle modules constitute the functional core of the Mediator complex38,39. Analysis of the promoter regions (1 kb upstream of the start codon) of all GmPR1 homologs revealed the presence of the as-1 (TGACG) element, a conserved binding site for TGA TFs (Supplementary Fig. 7a). Chromatin immunoprecipitation followed by qPCR (ChIP-qPCR) was employed to examine the recruitment of GmMED21 and GmMED6 to the GmPR1 promote in transgenic soybean hairy roots. The P1 (a sequence containing an as-1 element) region of the GmPR1-1 promoter exhibited robust enrichment of both GmMED21 and GmMED6. No significant enrichment of GmMED21 or GmMED6 was detected with the promoter of control gene, GmCYP2 (Supplementary Fig. 7b). These results suggest that GmMED21 and GmMED6 are integral to the transcription of SA-responsive genes, thereby positively regulating plant immunity. Furthermore, silencing of GmMED21 or GmMED6 via RNA interference (RNAi) significantly attenuated the transcriptional levels of GmPR1-1 and GmPR1-6 (Supplementary Fig. 7c). Functional validation demonstrated that the silencing of GmMED21 or GmMED6 in soybean root hairs led to a 1.7-fold increase in P. sojae oospore production and a 2-fold increase in pathogen biomass compared to the control, indicating markedly enhanced susceptibility (Supplementary Fig. 7d–f).
The interaction between MED21 and MED6 is crucial for the full activation of multiple activators40,41. In this study, we generated two GmMED21 truncation variants: a N-terminal deletion mutant (GmMED2116-139) and a discrete N-terminal fragment (GmMED211-15) (Fig. 4a). Y2H and LCI assays mapped the GmMED6 interaction with the N-terminal 15-aa domain of GmMED21. Notably, this same region mediates the interaction with PsAvh109. The GmMED211-15 retained the ability to interact with both PsAvh109 and MED6, whereas GmMED2116-139 failed to interact with either (Fig. 4b–d). These data indicate that the N-terminal 15 amino acids of GmMED21 constitute the shared binding interface for both proteins.
Fig. 4: PsAvh109 targets the N-terminal domain of MED21 to disrupt its interaction with MED6 and interfere with transcriptional regulation.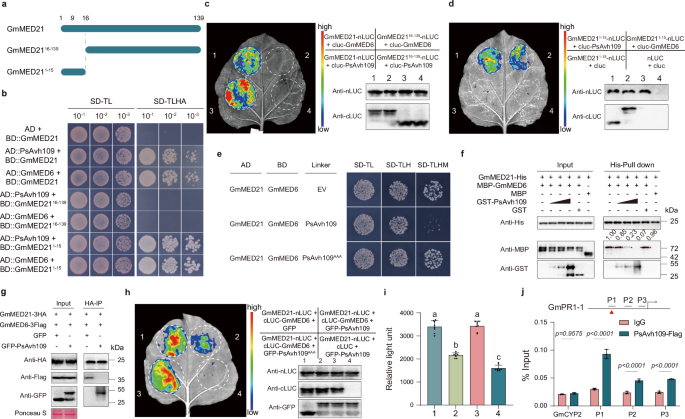 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a Schematic representation of full-length GmMED21 and truncated mutants: GmMED2116-139 (lacking the N-terminal 15 residues) and GmMED211-15 (N-terminal peptide only). b Yeast two-hybrid assay showing that full-length GmMED21 and GmMED211-15, but not GmMED2116-139, interact with PsAvh109 or GmMED6. Serial 10-fold dilutions were plated on SD-TL and SD-TLHA media; images captured after 4 days. c,d LCI in N. benthamiana confirms that the N-terminus (residues 1-15) of GmMED21 is necessary and sufficient for interaction with both PsAvh109 and GmMED6. Luminescence was detected 2 days post-Agrobacterium co-infiltration following D-luciferin application; protein expression was verified by Western blotting. e Yeast three-hybrid (Y3H) assay demonstrating that PsAvh109, but not PsAvh109AAA, disrupts the GmMED21-GmMED6 interaction. Growth on SD-TLHM (inducing PsAvh109 expression via methionine starvation) shows reduced interaction compared to empty vector or PsAvh109AAA controls. f In vitro competitive pull-down assay: increasing concentrations of GST-PsAvh109, but not GST, reduce MBP-GmMED6 binding to His-GmMED21 immobilized on Ni-NTA beads, indicating direct competition. Western blots probed with anti-GST, anti-MBP, and anti-His antibodies. g Co-immunoprecipitation in N. benthamiana: co-expression of GFP-PsAvh109, but not GFP, reduces GmMED6-3Flag co-precipitation with GmMED21-3HA using HA-Trap beads. Ponceau S staining served as loading control. h Luciferase complementation imaging shows that PsAvh109, but not PsAvh109AAA, suppresses GmMED21-GmMED6 interaction in planta; protein expression was verified by Western blotting. i Quantification of luciferase activity (Relative light unit, RLU) in treated leaves in (h). Data presented as mean ± SD (n = 8). Different letters indicate significant differences (P < 0.01; one-way ANOVA with Tukey’s multiple comparisons test). j Chromatin immunoprecipitation followed by qPCR (ChIP-qPCR) in PsAvh109-FLAG transgenic soybean hairy roots reveals enrichment of PsAvh109 at promoter regions (P1) containing as-1 cis-elements (red triangles) of immunity-related genes. IgG served as negative control; signals normalized to input DNA. Data: mean ± SD (n = 3; two-way ANOVA with Šídák’s test). (b–g) All the experiments were repeated three times with similar results.
LCI assays revealed a robust luminescence signal in the leaves of N. benthamiana upon co-infiltration of MED21 and MED6. However, when additional GmMED211-15-mCherry or full-length GmMED21-mCherry was expressed as a competitor, the luminescence intensity significantly decreased (Supplementary Fig. 8a, b). Similarly, GST pull-down assays indicated that GmMED211-15-mCherry competitively inhibited the interaction between GST-GmMED6 and GmMED21-His in a dose-dependent manner (Supplementary Fig. 8c). Functional assays showed that overexpression of GmMED211-15 in soybean hairy root significantly promoted P. sojae infection, leading to a 1.5-fold increase in oospore production and a 1.7-fold increase in pathogen biomass (Supplementary Fig. 8d–f). Compared with empty vector (mCherry) controls, RT-qPCR showed that the transcript levels of GmPR1-1 and GmPR1-6 were significantly down-regulated in MED211-15-overexpressing roots (Supplementary Fig. 8g). Collectively, these results indicate that disruption of the MED21-MED6 interaction suppresses of SA-responsive immunity.
Based on the shared binding domain, we hypothesized that PsAvh109 competitively displaces GmMED6 from GmMED21, thereby disrupting the host resistance. Yeast three-hybrid (Y3H) assays confirmed that PsAvh109, but not the PsAvh109AAA mutant, disrupted the GmMED21-GmMED6 interaction (Fig. 4e). Furthermore, in vitro Pull-down assays validated that GST-PsAvh109 reduced MBP-GmMED6 binding to GmMED21-His in a dose-dependent manner (Fig. 4f). Consistent with this, Co-IP assays revealed that PsAvh109-GFP co-expression attenuated the association between GmMED21-HA and GmMED6-3Flag (Fig. 4g). In LCI assays, co-expression of GFP-PsAvh109 with GmMED21-nLUC and cLUC-GmMED6 in N. benthamiana significantly reduced luminescence intensity compared to the GFP control, whereas GFP-PsAvh109AAA had no effect (Fig. 4h, i). ChIP-qPCR analysis revealed that PsAvh109 was significantly enriched at the P1 region of GmPR1-1 promoter (Fig. 4j). These findings suggest that PsAvh109 interferes with the assembly of the pre-initiation complex by outcompeting MED6 for MED21 binding, thereby modulating the transcription of SA-responsive genes.
The N-terminal domain of MED21 is a hotspot for competitive binding by TPL and PsAvh109
Due to the growth defense trade-off in plants, the transcription of immunity-related genes is generally inhibited during vegetative growth. In yeast, the general corepressor Tup1 hinders RNA polymerase II (Pol II) recruitment by outcompeting MED6 for binding to Srb7p (MED21); this interaction necessitates an intact Srb7p N-terminus24. Notably, the residues of MED21 that interact with Tup1 are identical to those required for the transcriptional repression by the Arabidopsis corepressor TOPLESS (TPL)24,25. Previous reports have shown that the TPD domain of TPL (TPLN188) is the primary determinant for the MED21 interaction22. We hypothesized that repressors TPL suppresses MED21-MED6 complex assembly by competing with MED6 for the MED21 N-terminal domain, and that PsAvh109 functionally mimics this mechanism to disrupt immune signaling.
Y3H assays demonstrated that the interaction between GmMED21 and GmMED6 was evidently inhibited in the presence of GmTPLN188 (Fig. 5a). Pull-down assays further confirmed that increasing concentrations of GST-GmTPLN188 reduced the abundance of MBP-GmMED6 associated with GmMED21-His (Fig. 5b). Co-IP assays showed that co-expressed GmTPLN188-GFP significantly attenuated the association between GmMED21-3HA and GmMED6-3Flag (Fig. 5c). LCI assays demonstrated that co-expression of FLAG-GmTPLN188 with GmMED21-nLUC and cLUC-GmMED6 significantly suppressed luminescence signals relative to GFP controls (Fig. 5d, e), suggesting that GmTPL suppresses the interaction between GmMED21 and GmMED6. ChIP-qPCR analysis revealed significant enrichment of GmTPL at the as-1 element (P1) of the GmPR1-1 promoter (Fig. 5f), indicating that GmTPL participates in the assembly of the pre-initiation complex site.
Fig. 5: GmTPL competes with GmMED6 for binding sites on GmMED21.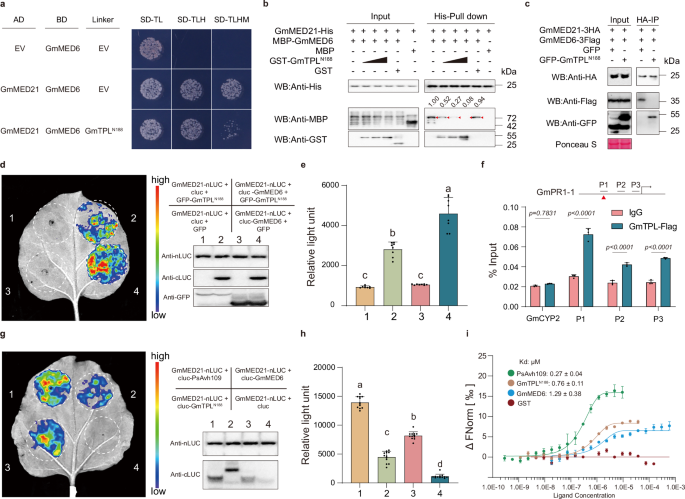 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a Y3H assay showing that expression of the N-terminal fragment of GmTPL (GmTPLN188) disrupts the interaction between GmMED21 and GmMED6. Yeast co-transformed with pGAD-GmMED21 and pBridge-GmMED6 were induced to express either empty vector (EV) or GmTPLN188 on SD-TLHM medium (inducing linker expression via methionine starvation). b In vitro competitive pull-down assay: increasing concentrations of GST-GmTPLN188, but not GST control, reduce MBP-GmMED6 binding to His-GmMED21 immobilized on Ni-NTA beads. Western blots probed with anti-GST, anti-MBP, and anti-His antibodies confirm dose-dependent competition. c Co-IP in N. benthamiana: co-expression of GFP-GmTPLN188, but not GFP, reduces GmMED6-3Flag co-precipitation with GmMED21-3HA using HA-Trap beads. Ponceau S staining served as loading control. d Luciferase complementation imaging shows that GmTPLN188 suppresses the GmMED21-GmMED6 interaction in planta; protein expression was verified by Western blotting. e Quantification of luciferase activity (Relative light unit, RLU) in treated leaves in (d). Data presented as mean ± SD (n = 8). Different letters indicate significant differences (P < 0.01; one-way ANOVA with Tukey’s multiple comparisons test). f ChIP-qPCR in GmTPL-FLAG transgenic soybean hairy roots reveal enrichment of GmTPL at promoter regions (P1) containing as-1 cis-elements (red triangles) of immunity genes. IgG served as negative control; signals normalized to input DNA (mean ± SD, n = 3; two-way ANOVA with Šídák’s test). g Comparative luciferase complementation assays show relative interaction strengths among PsAvh109, GmTPLN188, GmMED6, and GmMED21; protein expression was verified by Western blotting. h Quantification of luciferase activity in treated leaves in (g). Data presented as mean ± SD (n = 12). Different letters indicate significant differences (P < 0.01; one-way ANOVA with Tukey’s multiple comparisons test). i Microscale thermophoresis binding analysis using fluorescently labeled GmMED21 and titrated unlabeled partners reveals dissociation constants (Kd): PsAvh109 = 0.27 ± 0.04 μM, GmTPLN188 = 0.76 ± 0.11 μM, GmMED6 = 1.29 ± 0.38 μM; no binding detected for GST. (b,c,i) All the experiments were repeated three times with similar results.
Given that both Avh109 and TPLN188 compete with MED6 for binding to MED21, we compared the relative interaction affinities between GmMED21 and its respective partners (GmMED6, GmTPLN188, or PsAvh109). LCI assays revealed that the PsAvh109-GmMED21 interaction produced significantly higher luminescence intensity than the interactions of GmMED21 with either GmTPLN188 or GmMED6 (Fig. 5g, h). Furthermore, microscale thermophoresis (MST) measurements demonstrated that PsAvh109 binds GmMED21 with a dissociation constant (Kd) of 0.27 ± 0.04 μM, whereas GmTPLN188 and GmMED6 exhibited higher Kd values of 0.76 ± 0.11 μM and 1.29 ± 0.38 μM, respectively (Fig. 5i). These data indicate that PsAvh109 exhibits higher binding affinity to GmMED21 than GmMED6 or GmTPLN188. Therefore, PsAvh109 acts as a functional mimic of TPL, displacing TPL from MED21 to disrupt MED21-MED6 complex assembly and suppress immune signaling.
Both N- and C-termini of PsAvh109 are required for its interaction with GmMED21
Phylogenetic analysis revealed the conservation of PsAvh109 across Phytophthora species, including P. parasitica, P. infestans, and P. capsici. All PsAvh109 homologs featured NLS, RxLR-dEER, and TP motifs. Furthermore, the N-terminal 50 residues of PsAvh109 exhibit high sequence conservation among the homologs, whereas the C-terminal 15 residues consist of three invariant and three additional highly conserved residues (Supplementary Fig. 2b).
To further delineate the domains of PsAvh109 required for interaction with GmMED21, we generated a series of truncated mutants by deleting specific motifs or regions: the RxLR-dEER motif (designed as PsAvh109-RxLR), the TP motif (designed as PsAvh109-TP), the N-terminus of PsAvh109 (designed as PsAvh10951-167), the C-terminus of PsAvh109 (designed as PsAvh10921-152), or both termini (designed as PsAvh10951-152) (Supplementary Fig. 9a). Y2H assays revealed that three mutants, PsAvh10951-167, PsAvh10921-152 and PsAvh10951-152 lost their ability to interact with GmMED21, whereas the other truncated variants retained their binding capacity (Supplementary Fig. 9b). Consistently, LCI assays in N. benthamiana showed that the dual-terminal deletion mutant (PsAvh10951-152) produced no detectable luminescence. Conversely, only weak luminescence signals were observed for the single-terminal deletion mutants, PsAvh10951-167 and PsAvh10921-152 (Supplementary Fig. 9c). Additionally, all three mutants failed to inhibit INF1-induced PCD, similar to the PsAvh109AAA, whereas the wild-type PsAvh109 strongly inhibited PCD (Supplementary Fig. 9d). Collectively, these findings indicate that both the N- and C-termini of PsAvh109 are necessary for full interactions with GmMED21.
To identify the residues critical for the PsAvh109-GmMED21 interface, we conducted molecular modeling and protein-protein docking analysis using AlphaFold3. The predicted 3D structure of GmMED21 exhibited high structural homology with Arabidopsis thaliana MED21 (UniProt ID: C0LU16). Structural analysis revealed that specific residues within the N-terminal (V23, R40, Y41, V54, N58, Q66, Y69, F71) and C-terminal (W154, F157, W160, Y161, D167) of PsAvh109 contribute to MED21 binding. Together, they form a “headphone-like” structure that encircles the N-terminal region of MED21 (Supplementary Fig. 9e). This structural configuration likely accounts for the markedly higher binding affinity of PsAvh109 to MED21 compared to the native transcriptional repressor TPL. Notably, key interacting residues in the N-terminus (V23, R40) and C-terminus (F157, W160, D167) of PsAvh109 are highly conserved among its homologs (Supplementary Fig. 2b), supporting the evolutionary conservation of its function in targeting MED21.
Salicylic acid releases the binding between TPL and MED21 without impacting Avh109
To explore whether SA alleviates TPL-mediated transcriptional repression of SA-responsive genes, we examined TPL protein stability in N. benthamiana following transiently expressing and exogenous SA treatment. No significant difference in TPL accumulation was observed between SA-treated and EtOH control groups (Fig. 6a), thereby ruling out proteasomal degradation. Co-IP assays further demonstrated that exogenous SA significantly attenuated the association between GmMED21-3HA and GFP-GmTPLN188 compared to the EtOH control (Fig. 6b). In contrast, SA treatment had no discernible effect on the interaction between GmMED21-3HA and GFP-PsAvh109 relative to the EtOH control group (Fig. 6c). Additionally, LCI assays revealed that SA treatment reduced luminescence intensity driven by the GmMED21-nLUC and cLUC-GmTPLN188 interaction to approximately 70% of the level observed in the EtOH control (Supplementary Fig. 10). Conversely, SA treatment did not affect the luminescence resulting from the interaction between GmMED21-nLUC and cLUC-PsAvh109 (Supplementary Fig. 10). In ChIP-qPCR assays, SA treatment (100 μM) significantly reduced GmTPL enrichment at the as-1 sites (P1) of GmPR1-1 promoter; however, PsAvh109 occupancy remained unchanged (Fig. 6d). In summary, these findings demonstrate that SA mediates the disassembly of TPL-mediated blockade on MED21-MED6 interaction, whereas PsAvh109 competitively locks the MED21-MED6 interaction interface with higher affinity.
Fig. 6: Salicylic acid disrupts the interaction between TPL and MED21 without impacting the Interaction between Avh109 and MED21.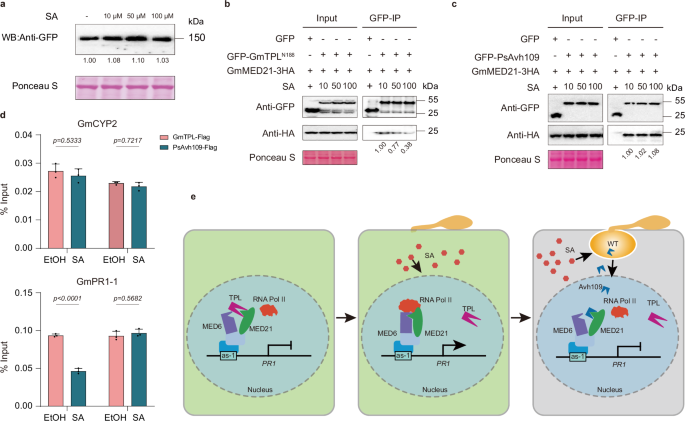 The alternative text for this image may have been generated using AI.
The alternative text for this image may have been generated using AI.
a Western blot analysis showing that exogenous SA treatment does not alter accumulation of GmTPL-GFP in N. benthamiana leaves. Ponceau S staining served as loading control. b Co-IP assay demonstrating that increasing concentrations of SA progressively reduce the interaction between GmTPLN188 and GmMED21. Anti-GFP beads were used to pull down GmTPLN188-GFP, and co-precipitated GmMED21 was detected by anti-HA immunoblotting. Ponceau S staining as a loading control. c Co-IP assay showing that SA treatment does not impair the interaction between PsAvh109 and GmMED21. GFP-PsAvh109 was immunoprecipitated, and associated GmMED21-HA levels remained unchanged across SA concentrations. Ponceau S staining confirmed equal loading. d ChIP-qPCR in transgenic soybean hairy roots expressing GmTPL-FLAG or PsAvh109-FLAG, treated with 100 μM SA or ethanol (control). SA treatment reduced GmTPL occupancy, but not PsAvh109 occupancy, at the P1 promoter region of GmPR1-1, which contains an as-1 element; GmCYP2 served as a negative control. Data are mean ± SD (n = 3; two-way ANOVA with Šídák’s test). e Model summarizing host-pathogen interplay at the Mediator complex: Under non-pathogen-infected conditions, GmTPL binds the N-terminus of GmMED21, preventing GmMED6 recruitment and repressing SA-responsive genes. Upon infection, SA accumulates and disrupts GmTPL-GmMED21 binding, enabling GmMED21-GmMED6 assembly and activation of immunity genes. However, P. sojae counters this defense by secreting PsAvh109, which retains binding to GmMED21 despite SA signaling, thereby blocking Mediator-mediated transcriptional reprogramming and suppressing immunity. (a,b,c) All the experiments were repeated three times with similar results.
Hence, we propose a model for this dynamic process (Fig. 6e): TPL constitutively represses SA-responsive genes by occupying the MED21-MED6 interface; upon SA accumulation, TPL is displaced, enabling Mediator assembly and defense activation. However, PsAvh109 is induced by host SA and competitively binds MED21 with higher affinity than TPL or MED6, perpetuating transcriptional repression.