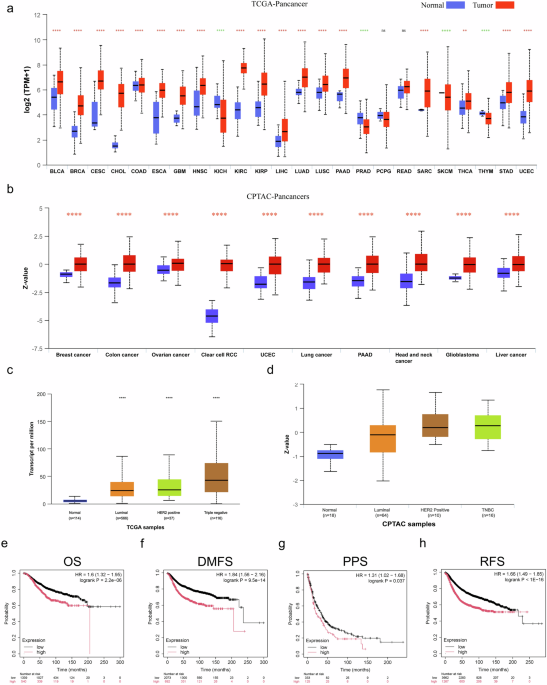
Expression and clinical correlation analysis of MCT4 (SLC16A3) across cancers
UALCAN (http://ualcan.path.uab.edu) is an open platform designed for user-friendly analysis of data from TCGA and CPTAC databases. We used UALCAN to conduct a pan-cancer analysis of SLC16A3 at both the transcriptional and protein levels, using TCGA and CPTAC data, respectively, with the gene specified as “SLC16A3” and the assay set to “Pan-Cancer Review.” To further investigate the expression of SLC16A3 in breast cancer, the cancer type was set to “breast cancer,” and the assay method to “expression assay.” Patients with breast carcinoma in TCGA and CPTAC were further stratified by stage and molecular subtype to assess differences in expression among these groups.
The Kaplan–Meier plotter (www.kmplot.com) is a web-based database that is used to assess the prognostic value of genes in various types of cancer. It integrates gene chip and RNA-Seq data from GEO, EGA, TCGA, and other public sources, enabling the evaluation of 70,632 genes across 21 cancer types, including breast (6234 cases), ovarian (2190 cases), lung (3452 cases), and gastric cancer (1440 cases). In this study, Kaplan–Meier survival curves were used to assess the prognostic value of MCT4 in breast cancer by analyzing its association with OS, DMFS, progression-free survival, and RFS. Hazard ratios with 95% confidence intervals and log-rank p-values were calculated to quantify the results.
Animals
All experimental procedures used in this study were approved by the Animal Care and Use Committee of Shenyang Pharmaceutical University (SYXK [Liao] 2021-0009). WT (MCT4+/+) BALB/c mice were purchased from Liaoning Changsheng Biotechnology Co, Ltd. under the animal license SCXK (Liao) 2020-0001. MCT4 homozygous knockout (MCT4-/-) BALB/c mice were developed by Beijing Biocytogen using CRISPR/Cas9-EGE technology, under the animal license SCXK (Jing) 2020-0007. SLC16A3 knockout alleles were generated by microinjection of in vitro-transcribed guide ribonucleic acids (gRNAs) containing the protospacer sequences 5′Guide #1 (5′-GACGCTGACCGCCTTGGGGA AGG) and 3′Guide #2 (5′-GCTTGACACCTTTACACACG GGG). PCR primers flanking the target site (WT-F 5′-GGCTGAGATGGTCACCCTGGCAACC; WT-F1 5′-TCCTGCTTGAGCCCAAAGTCACTG; WT-R 5′-GAGGGTATAAGGAGAGCAGATAGGG; WT-R1 5′-TGGGGTGTCAGGTCCTTAGTGCAGG) and a primer within the knockout sequence (Mut-R: 5’-GAGGGTATAAGGAGAGCAGATAGGG) were used to amplify and identify the modified alleles.
Mice were housed in individually ventilated cages under the specific pathogen-free facility of Shenyang Pharmaceutical University. They were housed in groups on a 12-h light/dark cycle with ad libitum access to food and water at 20–22 °C. All experiments were performed during the day cycle. Unless otherwise specified, both male and female mice were included in the analysis, and all mice were 5–6 weeks old. Mice were randomly assigned to groups.
Cell culture
The 4T1 cells were obtained from the American Type Culture Collection (Manassas, VA, USA). Stable 4T1 cells overexpressing MCT4 (4T1-MCT4) were generated via plasmid transfection by electroporation, as previously described20. The 4T1-Luc cells were purchased from Hunan Fenghui Biotechnology Co., Ltd. (Hunan, China). Professor Cong Luo (Shenyang Pharmaceutical University) provided RAW264.7 cells. All cell lines were cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% fetal bovine serum and 1% penicillin–streptomycin. M109 cells obtained from Shanghai Guan&Dao Biological Engineering Co., Ltd. (Shanghai, China) were cultured in RPMI 1640 medium containing 10% fetal bovine serum and 1% penicillin–streptomycin. All cell lines were maintained at 37 °C in a humidified atmosphere with 5% CO2.
Food and water intake and body weight monitoring
WT and MCT4-/- mice (4 weeks old; seven males and seven females) were used in the experiments. The weekly food and water intake was recorded to calculate the daily consumption per mouse, and body weight changes were monitored to assess the effect of MCT4 knockout on BALB/c mice.
Glucose tolerance test
Three WT and MCT4-/- mice of each sex were randomly selected to undergo a 12-h fasting period before participating in the intraperitoneal glucose tolerance test. A bolus of glucose (2 g/kg) was injected intraperitoneally. The blood glucose levels of each mouse were measured at 0, 15, 30, 60, and 120 min.
Tumor allograft model
To establish 4T1 and M109 tumor models, 3 × 106 cells (0.1 mL) were subcutaneously injected into the right upper abdomen of 6-week-old WT and MCT4-/- mice (four males and four females per genotype). The long and short diameters of the tumor were recorded every 7 days using digital calipers, and the tumor size was determined using the formula (width2 × length)/2. In growth curve studies, the animals were sacrificed 21 days after implantation when the tumor was growing exponentially. Female mice were transplanted with 4T1-Luc cells using a consistent protocol, followed by in vivo imaging and quantification of average fluorescence intensity to assess differences in tumor growth. To document the survival duration of tumor-bearing mice, the animals were monitored daily over 40 days, with survival analysis conducted at the conclusion of the observation period.
The effect of IGF1 and lactate on tumor growth was assessed using 4T1 allograft models. Briefly, IGF1 or lactate was administered to MCT4-/- mice via subcutaneous injection at a dosage of respectively 3 and 15 mg/kg on days 7, 10, 13, 16, 19, and 22 after engraftment. The tumor size was determined using the formula (width2 × length)/2; mice were euthanized when the tumors were ulcerated or when the tumor volume reached 1500 mm3.
Breast cancer-based lung metastasis model
To generate a breast cancer mouse model, 4T1-Luc cells (1 × 106 cells/mouse) were resuspended in phosphate-buffered saline (PBS) and slowly injected into the tail vein of 6-week-old female WT and MCT4-/- mice divided into two groups of four. Three weeks after injection, the mice were anesthetized with isoflurane and administered D-fluorescein potassium salt. Subsequently, in vivo imaging was performed, and the mean fluorescence values of the lung metastases were quantified.
Protein extraction and western blotting
Cells or tissues were lysed in ice-cold RIPA buffer containing a protease inhibitor cocktail. The samples were centrifuged at 12,000 × g for 15 min at 4 °C. The supernatant was collected, and the protein concentration was determined using the BCA Protein Assay kit (Cat. No. AR0197A, Wuhan Boster Biological Technology, Ltd.).
To perform western blot analysis, lysates (15–50 µg/lane) were resolved using sodium dodecyl sulfate-polyacrylamide gel electrophoresis with a 12% gel. The resolved proteins were transferred to a 0.45-µm polyvinylidene difluoride membrane (Cat. No. IPFL00010; Merck KGaA). Subsequently, the membrane was blocked with 5% bovine serum albumin (BSA) in Tris-buffered saline containing 0.1% Tween 20 (TBST) for 1 h at room temperature to reduce background staining. The membranes were then incubated overnight at 4 °C with primary antibodies (anti-MCT4, Cat. No. 22787-1-AP; anti-beta actin, Cat. No. 66009-1-Ig; Proteintech). After washing with TBST, horseradish peroxidase–conjugated secondary antibodies (anti-mouse, Cat. No. 31430; anti-rabbit, Cat. No. 31460; Thermo Fisher Scientific, Inc.) were applied for 1 h at room temperature. The protein bands were visualized using enhanced chemiluminescence (Cat. No. KGP1127, Nanjing KeyGen Biotech Co., Ltd.) with 5200 Multi Automatic Chemiluminescence (Tanon Science and Technology Co., Ltd.).
TIF isolation
TIF was isolated from tumors using a previously described centrifugal method53. Tumor-bearing animals were euthanized via cervical dislocation, and the tumors were rapidly excised. Following dissection, the tumors were weighed, rinsed briefly in saline (150 mM NaCl) at room temperature, and blotted on filter paper. An equivalent mass of tumor tissue was placed into 1.5-mL conical tubes and centrifuged at 10,000 × g for 10 min at 4 °C. TIF was collected from the conical tube and stored at −80 °C until further analysis.
Measurement of lactate and IGF1
The lactate concentration in the culture supernatant, serum, and TIF was measured using the L-lactate detection kit (Cat. No. A09-2-1, Nanjing Jiancheng Bio-Engineering Institute) following the manufacturer’s protocol. A Mouse IGF1 ELISA Kit (Cat. No. EK0378, Wuhan Boster Bio-Engineering Co., Ltd) was used to quantify the IGF1 concentration in these samples using a double-sandwich method.
Analysis of lymphocyte subsets
The populations of CD4+ T cells, CD8+ T cells, NK cells, and macrophages were analyzed using fluorescence-activated cell sorting. For immune cell population analysis in peripheral blood, approximately 300 µL of blood was collected from the retro-orbital sinus of mice into heparinized tubes. Peripheral blood mononuclear cells (PBMCs) were isolated using the mouse peripheral blood lymphocyte isolation kit (Cat. No. C009, Beyotime) following the manufacturer’s protocol. The staining protocol for peripheral blood lymphocyte subsets was as follows: anti-CD45-FITC/CD3-APC/CD4-PE antibody for labeling CD4+ T cells; anti-CD45-FITC/CD3-APC/CD8-PE antibody for labeling CD8+ T cells; anti-CD45-FITC/CD3-APC/CD49b-PE antibody for labeling NK lymphocytes; anti-CD45-FITC/CD11b-APC/F4/80-PE antibody for labeling macrophages; anti-F4/80-PE/CD80-FITC antibody for labeling M1-polarized macrophages; anti-F4/80-PE/CD206-FITC antibody for labeling M2-polarized macrophages; and anti-CD45-FITC/F4/80-PE/ Ly6G Ly6C-APC antibody for labeling MDSCs. The antibodies used in this study are listed in Supplementary Table 2. The stained cells were washed, resuspended in 0.1% BSA-containing PBS, and analyzed using a BD FACSAria™ III flow cytometer (BD Biosciences). To analyze immune cell populations in fresh tumors, single cells were isolated using collagenase IV and stained, and the immune cell populations in peripheral blood were analyzed using the same methodology.
In vitro macrophage polarization experiments
The effect of MCT4 on macrophage polarization was investigated using RAW264.7 cells. The cells were adjusted to a concentration of 1 × 106 cells/mL in Dulbecco’s modified Eagle’s medium and treated with IL-4 (100 ng/mL) to induce M2 polarization. Subsequently, the cells were cultured for 24 h with 4T1 culture supernatant, 4T1-MCT4 culture supernatant, and 4T1 culture supernatant containing IGF1 (300 ng/mL)/lactate (10 mM) for 24 h. Anti-mouse CD206-APC antibody was used to assess M2 polarization. The stained cells were washed, resuspended in 0.1% BSA-containing PBS, and analyzed using a BD FACSAria™ III flow cytometer (BD Biosciences).
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from 4T1 and 4T1-MCT4 cells using TRIzol Universal Reagent (Cat No. DP424, Tiangen Biotech Co., Ltd.). Complementary DNA was synthesized from the RNA using the HiFiScript cDNA Synthesis Kit (Cat No. CW2569, CoWin Biosciences) under the following conditions: 42 °C for 15 min and 85 °C for 5 min. RT-qPCR analysis was then performed using TB Green® Premix Ex Taq™ II (Cat No. RR820A, TAKARA) on a CFX96 system (Bio-Rad Laboratories, Inc.) with the appropriate primers (Supplementary Table 3). The RT-qPCR conditions were as follows: initial activation at 95 °C for 3 s, followed by 40 cycles of denaturation at 95 °C for 5 s and annealing at 60 °C for 30 s. mRNA expression levels were calculated using the 2−ΔΔCt relative quantification method54. The expression level of each target gene was normalized to that of β-actin mRNA in the same sample.
Breast, lung, and oral carcinogenesis induction mouse model
WT and MCT4-/- mice (6 weeks old) were used to establish models for tumor carcinogenesis induction. In the breast cancer induction model55, mice were injected subcutaneously with 100 μL of DMBA (2 mg/mL) every 2 weeks over a total duration of 20 weeks, followed by 8 weeks of plain water supplementation. In the lung cancer induction model56, mice received intraperitoneal injections of 100 μL of urethane (100 mg/mL) weekly for 18 weeks, followed by plain water supplementation for 7 weeks. In the oral carcinogenesis induction model57, mice were fed 4NQO at a final concentration of 50 μg/mL in their drinking water for 22 weeks, followed by 7 weeks of plain water supplementation. The lesions were examined weekly, and their weight was monitored as an indicator of pain. Mice were sacrificed at the end of the experiment.
Hematoxylin and eosin staining
Fresh tumor tissues were fixed in 4% paraformaldehyde for 24 h, followed by gradient alcohol dehydration and embedding in paraffin. Sections (5 μm) were dewaxed in xylene, rehydrated using graded alcohol, and heat-treated at 62 °C for 2 h. After antigen retrieval, the slides were stained with hematoxylin, differentiated in 1% acid alcohol, and counterstained with 5% eosin. Dehydration was performed using graded alcohol (80%, 95%, and 100%) and xylene, followed by neutral gum mounting. A Nikon ECLIPSE-E200 microscope was used for histological analysis; three non-overlapping cortical fields were randomly selected for imaging.
Immunofluorescence (IF)
Fresh tumor tissues were fixed in a 4% polymethyl aldehyde solution for 24 h. All samples were paraffin-embedded and sectioned. Briefly, 5-μm tumor sections were deparaffinized, subjected to antigen retrieval, blocked, and stained with primary antibodies. Subsequently, the sections were stained with fluorescence-conjugated secondary antibodies, and the nuclei were stained with DAPI. Random observations of the tumor tissue cortex were performed at 400× magnification using a fluorescence microscope (NIM900 Ningbo Yongxin Optics Co., Ltd.).
Statistical analysis
Flow cytometry data were analyzed using FlowJo 10.8.1 software. Data representation, statistical testing, and outlier identification were performed using GraphPad Prism 10.1.2. The results are represented as the mean ± standard error of the mean. Two-tailed unpaired t-tests were used to compare results between two groups. One- or two-way ANOVA with Tukey’s post hoc test was used to compare results among more than two groups. Correlation was assessed by Pearson’s coefficient. P-values < 0.05 were considered to indicate a statistically significant difference. Exact p-values and biological replicates (N) are reported in the figure legends.
Ethics statement
All animal experiments performed in this study were reviewed and approved by the Animal Care and Use Committee of Shenyang Pharmaceutical University (Approval Number: SYXK [Liao] 2021-0009). This study was conducted in strict compliance with the institutional guidelines for the care and use of laboratory animals. We have complied with all relevant ethical regulations for animal use.
Animal Welfare and Euthanasia: All efforts were made to minimize animal suffering and distress. Mice were housed in a specific pathogen-free (SPF) facility under a 12-hour light/dark cycle with ad libitum access to food and water. At the experimental endpoints, animals were euthanized humanely. The primary method of euthanasia for tissue collection was cervical dislocation under deep anesthesia induced by isoflurane inhalation (5% for induction, 1.5–2% for maintenance). For in vivo imaging procedures requiring terminal anesthesia, euthanasia was ensured by an overdose of isoflurane followed by cervical dislocation as a secondary physical method. All procedures were performed by trained personnel.
Tumor Burden Monitoring: The maximal tumor volume permitted by our approved protocol was 1500 mm³. Tumor size was measured every 2-3 days using a digital caliper, and volume was calculated using the formula: (width² × length)/2. Animals were monitored daily for signs of distress (e.g., lethargy, hunched posture, difficulty moving, or ulceration). In no instance throughout this study was the maximal permitted tumor volume exceeded. Any animal showing signs of significant discomfort or reaching the predefined humane endpoint (tumor volume ≥ 1500 mm³ or ulceration) was euthanized immediately and excluded from subsequent longitudinal analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.