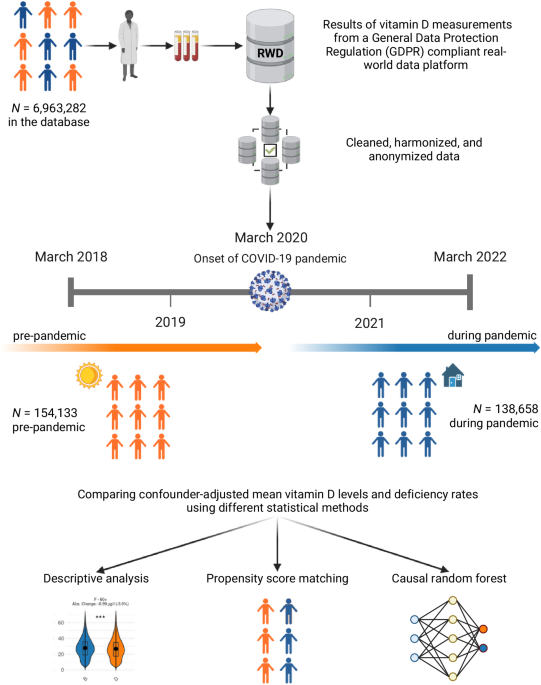
Study design
This retrospective analysis used routinely-collected laboratory data to analyze vitamin D levels in the Bavarian population before and during the COVID-19 pandemic. The primary objective was to evaluate population-wide changes in vitamin D levels by comparing the pandemic period as defined by the WHO19 with a pre-pandemic period of equal duration. Specifically, the pre-pandemic period was defined as March 2018 to February 2020, and the pandemic period was defined as March 2020 to February 2022. Both time periods were of equal length to ensure comparability. Our analysis focused on the overall, population-wide impact of the COVID-19 pandemic on vitamin D levels, rather than examining individuals’ COVID-19 infection status.
Data source
Data on vitamin D levels were extracted from the Honic data platform12, which provides quality-controlled, routinely-collected healthcare data from multiple sources, including laboratory reports, outpatient visits, and prescriptions. For the current study, data primarily originated from a large laboratory chain within Bavaria comprising over 10 laboratories. Data is transmitted in real time from laboratory information systems to the Honic data platform, with approximately 12,000 to 15,000 new reports added daily. The database encompasses a highly diverse population, including individuals from both inpatient and outpatient settings, all types of health insurance, and a wide range of practitioner types, such as general practitioners, specialists, and hospital-based physicians. At the time of data extraction (03/2024), the database included over 7 million individuals, providing a comprehensive representation of real-world healthcare interactions in Bavaria.
Through the Honic platform, RWD is collected, cleaned, processed, and standardized to ensure high-quality, reliable data for analysis. The platform uses a master patient index to link patient data across sources while ensuring privacy through pseudonymization. Data access is provided within a secure analytics environment, ensuring compliance with the General Data Protection Regulation in the European Union and other data protection regulations.
Data extraction
All available 25-hydroxy vitamin D measurements taken during the defined time frames were extracted from the Honic data platform for this study. To maintain consistency, only the first available vitamin D measurement per individual within the study period was included, focusing on baseline levels and minimizing potential confounding due to treatment or seasonal variations from repeat measurements. Additional variables such as age, gender, postal codes, and season were also extracted.
Inclusion criteria
Individuals were required to have at least one vitamin D measurement within the specified time frames, as well as the following criteria: (1) Age: Adults 18 years and older at the time of vitamin D measurement. (2) Residence: Individuals residing in Bavaria, defined by postal codes beginning with 60 or ranging from 80 to 97, to ensure a regionally homogeneous sample. (3) Administrative gender: Male or female individuals. (4) Vitamin D measurement: Individuals with at least one valid value. Figure 2 shows a detailed overview of the selection criteria for the study population.
Data processing
Prior to analysis, the extracted data was processed and anonymized to ensure data security and regulatory compliance. For this, variables that were redundant for the purpose of our analysis were removed, age was aggregated into age groups to reduce the risk of re-identification, postal codes were aggregated to degree of urbanization, and any specific identifiers were removed. As part of the latter step, postal codes (after applying the selection criteria), precise dates (retaining only month and year), and extreme outliers (that could lead to re-identification) were removed. A re-identification risk assessment was conducted on the remaining attributes, and k-anonymity (with k = 5) was applied to quasi-identifiers. Anonymization was performed using ARX version 3.9.144. The final dataset was formatted as a CSV file and transferred to a secure R server for analysis.
Measures
Vitamin D was assessed through serum levels of 25-hydroxy vitamin D (25-OH-D), the standard biomarker for evaluating vitamin D status in clinical and research settings. This biomarker is preferred because it reflects both dietary intake and endogenous production from sunlight exposure. Serum 25-OH-D concentrations were measured using immunoassays, a widely used method in routine laboratory diagnostics18,45. Vitamin D levels below 4 μg/L or above 300 μg/L were recorded as “<4 μg/L” or “>300 μg/L”, respectively, representing the assay’s detection limits. Since exact values could not be extrapolated for these cases, individuals with such recordings were excluded from the mean vitamin D levels analyses (0.4% excluded) but retained for deficiency rate calculations. According to widely accepted guidelines, a serum 25-OH-D level below 20 μg/l (50 nmol/l) was defined as vitamin D deficiency6,46. Outliers (values > 70 μg/l, corresponding to 0.96% of the data) were removed only for plotting to enhance visualization and prevent extreme values from distorting the scales. However, all statistical analyses, including group means and significance tests, were conducted on the full dataset to ensure the results remain representative of the population. The seasonal patterns of vitamin D levels over time are shown in Supplementary Fig. S1. A comparison between vitamin D levels during early vs. late stages of the pandemic further revealed no substantial difference (Supplementary Fig. S2).
We further use the following covariates to account for different sources of heterogeneity. (1) Age: The age of individuals was calculated at the measurement date and pre-grouped into three categories: 18–39 years, 40–59 years, and 60+ years. (2) Gender: The administrative gender was coded as male or female, individuals with unknown gender were excluded from the analysis. (3) Season: Seasons were categorized as follows: winter (December, January, and February), spring (March, April, and May), summer (June, July, and August), and fall (September, October, November).
Statistical analysis
For comparisons of continuous variables between groups (e.g., vitamin D levels before and during the pandemic), we applied two-sided Wilcoxon rank-sum tests (wilcox.test in R) when normality assumptions were not satisfied, and two-sided t-tests (t.test) otherwise. Comparisons of categorical variables (e.g., gender or age group distributions) were performed using two-sided χ2-tests (chisq.test). All null hypothesis tests were conducted two-sided. Effect sizes from regression models are reported as standardized or unstandardized regression coefficients (β) with corresponding confidence intervals, p values, and degrees of freedom where applicable. Degrees of freedom for t- and F-statistics correspond to the residual degrees of freedom from the respective models. Confidence intervals were reported at the 99% level. A significance threshold of α = 0.01 was applied throughout our analysis. The statistical tests allow for comparisons at the population level but do not account for potential differences in individual characteristics between the periods. Therefore, we employed two additional methods to make comparisons: (1) propensity score matching and (2) a causal forest model. These methods were specifically designed to reduce bias arising from differences in individual characteristics across the two populations and thus yield confounding-adjusted estimates.
Propensity score matching
PSM20 was performed using a logistic regression model to estimate propensity scores, with the combination of age group, gender, and test month included as covariates. Nearest neighbor matching was applied, ensuring exact matches for gender, age group, and test month to create comparable pre-pandemic and pandemic groups. Test month was chosen as a covariate instead of season because it provides finer granularity in capturing potential seasonal effects and better aligns with the temporal distribution of laboratory data. After matching, the original sample of 292,187 observations was reduced to 266,820 matched observations, resulting in a loss of 25,367 records (8.7%). This matching procedure achieved strong balance across most covariates, with SMDs falling below the recommended threshold of 0.1 for the majority of variables. Two covariates exhibited slightly higher SMDs (−0.1200 for the test month of April and 0.1024 for the age group 18–39), which are marginal deviations from the threshold. These slight imbalances are unlikely to substantially affect the comparability of the matched groups47. By matching study populations in this manner, inferences can be drawn by comparing the intervention group (i.e., individuals tested during the pandemic) with the matched control group (i.e., individuals tested before the pandemic)48. See Supplementary Table S2 for more details on the population characteristics after matching. After PSM, adjusted comparisons were conducted directly within the matched sample without fitting an additional regression model. Specifically, we compared mean vitamin D levels between periods using independent sample t-tests and compared vitamin D deficiency rates using χ2-tests.
Causal forest
We further employed the causal forest model, a nonparametric state-of-the-art machine learning method13,14,49. The causal forest aims at estimating heterogeneous treatment effects and is thus well-suited for our task. First, causal forests share many favorable properties with random forests in that they are expressive, require little tuning, and have a low risk of overfitting, which often leads to a robust performance in practice. Second, under common mathematical assumptions, causal forests have been shown to be pointwise consistent for the true treatment effect and therefore lead to provably valid inferences14. Third, the causal forest makes few parametric assumptions of the form of the heterogeneous treatment effect and can thus model non-linear relationships, even in the presence of high-dimensional covariates, and typically offers better statistical power than other estimators.
The causal forest extends the traditional random forest50 by estimating changes in an outcome variable due to an intervention, while controlling for other observed confounders (here: age, gender, and season). The causal forest proceeds by first building an ensemble of decision trees to model the relationship between intervention, outcome, and covariates. For each tree, it splits the data recursively into subgroups based on covariates, so that heterogeneity in the change of the outcome variable due to the intervention is discovered. To avoid overfitting and ensure unbiased estimates, causal forests use a technique called “honesty”14, where one part of the data is used to determine the splits (i.e., how to group similar individuals) and where another part of the data is used for estimating the changes in the outcome variable.
The causal forest model was estimated for both outcomes of interest, namely, changes in vitamin D levels and vitamin D deficiency rates. The following covariates were used: age, gender, and season. Here, season was chosen over test month in the machine learning analysis as it captures broader environmental and behavioral patterns influencing vitamin D levels, while reducing potential noise from month-to-month variations. The causal forest model was run with the default parameter of 2000 trees51, ensuring sufficient depth and robustness in estimating heterogeneous treatment effects. All other hyperparameters were set to their default values as in ref. 51.
To validate the ignorability assumption, we applied balance diagnostics by checking the weighted absolute standardized mean difference (ASMD) for the three key covariates (age group, gender, and test month) between the pre-pandemic and pandemic study populations. A value close to 0 indicates that the propensity scores are well-calibrated. In our analysis, all ASMD values were well below the recommended 0.1 threshold, confirming a strong balance between groups. For further details, see Supplementary Table S5.
Comparison of statistical methods
To systematically compare the estimated confounder-adjusted reductions, we proceeded as follows. We use Cohen’s d52 for descriptive statistics, the SMD53 for PSM, and the average treatment effect as provided by causal forest. Since Cohen’s d and SMD are standardized measures, while the ATE is not, we re-transformed Cohen’s d and SMD into the same scale as the ATE to allow for comparability.
Subgroup analysis
To understand the variability in vitamin D levels across different subpopulations, we performed a subgroup analysis by evaluating the estimated changes within specific subgroups defined by age, gender, and season. First, we categorized the individuals into predefined subgroups: three age groups (i.e., 18–39, 40–59, 60+), two gender groups (i.e., women, men), and four season groups (i.e., winter, spring, summer, fall). We then ran a separate causal forest model for each subgroup. For each subgroup, we reported the estimated confounder-adjusted reduction in the outcome variable together with 99% and 95% confidence intervals (CI) and the percentage change relative to the average value across all individuals.
Robustness checks
To verify the robustness of our findings, we performed a series of robustness checks. First, we tested alternative time frames for the pre-pandemic and pandemic periods. The original analysis (March 2018–February 2020 vs. March 2020–February 2022) showed a mean vitamin D reduction of −0.7 μg/l and a deficiency rate increase of +4.0% (descriptive), with PSM-adjusted values of −0.5 μg/l and +3.6%. Adjusting the pre-pandemic period to end in January 2020 (i.e., March 2018–January 2020) and including February 2020 in the pandemic period yielded only a slightly larger difference (−0.9 μg/l, +4.9%), while PSM estimates remained unchanged (−0.5 μg/l, +3.6%). Also, adjusting the pre-pandemic period to include March 2020 (i.e., March 2018–March 2020) resulted in consistent findings (−0.6 μg/l, +3.2%; PSM: −0.5 μg/l, +3.3%).
Second, we restricted the analysis to the months of March, April, and May in 2018, 2019, 2020, and 2021. This analysis confirmed the reduction in mean vitamin D levels (−2.3 μg/l) and the increase in deficiency rates (+4.3%). This is further corroborated by the PSM-adjusted estimates, which compute to −1.6 μg/l and +6.2%, respectively. When restricting the data to the above-defined months, the causal forest estimated a reduction of −2.042 μg/l, corresponding to an − 8.1% decline during the pandemic compared to the pre-pandemic period. Together, the checks help rule out that the reductions rely on the specific choice of a month/season.
Third, we incorporated hours of sunshine (taken from54) as an additional covariate. Note that, due to the variation in the sunshine hours at the monthly level, it was not possible to perform PSM with exact matching, and, instead, we focused on the causal forest. The confounder-adjusted model estimated a reduction of −6.45 μg/l, indicating an even stronger decreasing in vitamin D levels during the pandemic, showing that the observed decline was not solely driven by seasonal sunlight variations.
Fourth, to test for unmeasured confounding, we conducted a negative control outcome analysis55, which involves selecting an outcome that should not be affected by the intervention, given proper adjustment for confounders. Due to the absence of other variables that could act as such interventions, we randomly shuffled the outcome variable (i.e., vitamin D values) within the dataset and then repeated all analyses. As expected, this yielded no meaningful differences between the pre-pandemic and pandemic periods (descriptive: −0.1 μg/l, +0.1%; PSM: −0.1 μg/l, +0.1%; ML: ATE −0.0781 μg/l). Hence, the results reveal no significant association between intervention and negative outcome, indicating that unmeasured confounding was unlikely to bias the findings.
Software
All statistical analyses were performed using the statistical software R (version 4.4.1)56. For PSM, we use the matchit function from the package MatchIt (version 2.3.2)57. For the causal forest, we used the package grf (version 4.5.5)51. For the sensitivity analysis, we used the package sensemakr (version 0.1.6)58.
Ethics approval
The data request underwent a comprehensive assessment by an external compliance board, including patient and scientific representatives, to ensure the project met the ethical and legal standards required for using Honic data. This study received an Institutional Review Board exemption from the Ethics Committee of the Medical Faculty of LMU Munich (4-0900-KB).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.