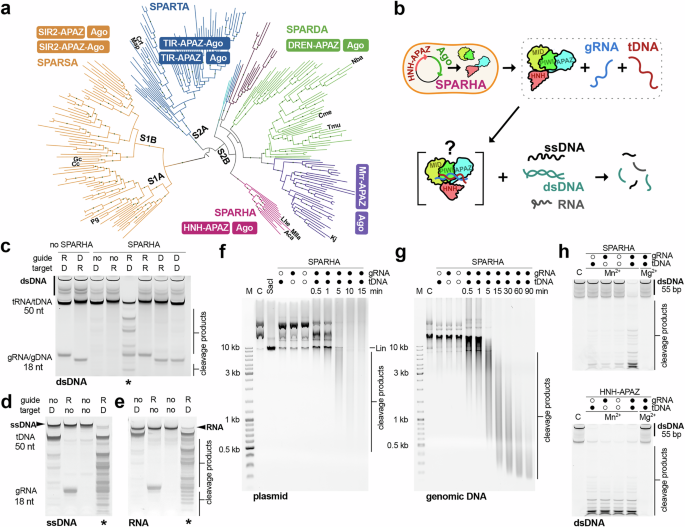
Bioinformatic analysis of short pAgos and HNH nucleases
For the construction of the phylogenetic tree of APAZ-containing proteins, their search was performed using DELTA-BLAST (part of the BLAST 2.15.0+ package) against the nr (non-redundant proteins) RefSeq database with the following parameters: -taxids 2,2157 (Bacteria, Archaea), -num_iterations 3, -evalue 0.005. The input file contained the APAZ domains of the following proteins: WP_141644551.1, WP_215874058.1, WP_075600999.1, WP_033317603.1, WP_011516871.1, WP_046756489.1, WP_010942011.1, WP_235024190.1, WP_006053074.1, WP_109649956.1, WP_092459739.1. The identified proteins were filtered using a custom Python script, resulting in 1546 unique proteins with WP identifiers. Their sequences, as well as the sequences of neighboring proteins in the genomes, were obtained using another custom Python script via Entrez Efetch and were then annotated using rpsblast (part of the BLAST 2.15.0+ package) and InterProScan 5.72-103.0. Protein sequences for which neighboring proteins containing the PIWI domain were extracted for further analysis and clustered using MMseq2 v.16.747c6 with a sequence identity threshold of 45%. After clustering, the previously studied proteins and the proteins from this study (identifiers listed above) were added to the resulting sequences, and a total of 321 sequences were aligned using MAFFT v.7.526 (method E-INS-i). The alignments were then trimmed with trimAl v.1.5 using the parameters -gt 0.5 (gap threshold), -resoverlap 0.65, -seqoverlap 65, resulting in 272 sequences. The phylogenetic tree was constructed using IQ-TREE v.2.3.6 with the ModelFinder algorithm (which identified Q.pfam+F + R7 as the best-fit model) and ultrafast bootstrap with 1000 replicates, followed by visualization using iTOL.
For the construction of the phylogenetic tree of HNH domain-containing proteins, a search was performed using DELTA-BLAST (part of the BLAST 2.15.0+ package) against the nr (non-redundant proteins) RefSeq database with the following parameters: -taxids 22157 (Bacteria, Archaea), -num_iterations 5, -evalue 0.005. The input file contained the sequences of HNH domains from various defense systems (Septu, Retron, Cas9, IscB, Zorya II, CRISPR I-F, EVE, SRA, Cap5 CBASS, Vvn, homing endonuclease). The identified proteins were filtered as described above, resulting in 8412 unique proteins with WP identifiers. Their sequences were obtained and clustered as described above. After clustering, the previously studied HNH domain-containing proteins (identifiers listed above) were added to the resulting sequences, and a total of 727 sequences were aligned as described above and trimmed with trimAl v.1.5 (-gt 0.5), resulting in 724 sequences. The phylogenetic tree was constructed and visualized as described above (the selected best-fit model was VT + I + R9). Protein annotation was performed using RPSBLAST, InterProScan 5.72-103.0, HHsuite v. 3.3.0 with the Pfam-A database, AlphaFold 3 (AF3) and the DALI server, as well as analysis of their genomic neighbors.
Cloning, expression, and purification of SPARHA
The coding sequences for AcaAgo, MllaAgo, LheAgo, and their partner proteins HNH-APAZ were codon-optimized for E. coli expression, obtained by custom gene synthesis and cloned into the pBAD-HisB expression plasmid downstream of the araBAD promoter with a ribosome binding site between each pair (pAgo and HNH-APAZ), resulting in the pBAD_AcaSPARHA, pBAD_MllaSPARHA, and pBAD_LheSPARHA plasmids. Site-directed mutagenesis was performed to obtain deletions and alanine substitutions in the sensor loop (Δ290-296 in AcaSPARHA, Δ291-297 in MllaSPARHA), PIWI-PIWI interface (quadruple substitution in MllaSPARHA including R273A, D275A in the interface loop and Q320A, R349A in the connector loop; correspond to Q272, N274 and Q, D in AcaSPARHA), the active site of the HNH domain (H53A in MllaSPARHA and N71A/H80A in AcaSPARHA), and in the N-terminal α helix and the bridge interface of the HNH domain (Q15A/K16A and E50A, respectively, in MllaSPARHA). For protein expression, E. coli BL21(DE3) cells harboring pBAD_SPARHA plasmids were grown in LB medium with ampicillin (150 μg/ml) at 37 °C with shaking (200 rpm) to OD600~0.5, followed by cold shock at 0 °C for 45 min. Protein expression was induced with 0.05% L-arabinose and carried out overnight at 16 °C. Then, the cells were harvested by centrifugation and used for protein purification.
To purify the SPARHA complexes, cells were lysed by sonication in buffer containing 20 mM HEPES, pH 8.0, 1 M KCl, 10 mM K2HPO4, 5% glycerol, 2 mM DTT, 2 mM PMSF and then centrifuged at 25,500 × g for 25 min at 4 °C. The supernatant was transferred onto a HisTrap HP 1 ml column (Cytiva) pre-equilibrated with buffer containing 20 mM HEPES, pH 8.0, 0.5 M KCl, 5% glycerol, washed with the same buffer and then with the buffer containing 21 mM imidazole, and the proteins were eluted with the buffer containing 300 mM imidazole. The eluate was supplemented with EDTA (up to 5 mM) and diluted 20-fold with buffer containing 20 mM HEPES, pH 8.0, 5% glycerol, 2 mM EDTA, and loaded onto a HiTrap Heparin HP column 5 ml (Cytiva) pre-equilibrated with buffer containing 20 mM HEPES, pH 8.0, 30 mM KCl, 5% glycerol and 2 mM EDTA. After washing, the complexes were eluted with a KCl gradient 30 to 800 mM in buffer containing 20 mM HEPES, pH 8.0, 5% glycerol, and 2 mM EDTA. For AcaSPARHA and LheSPARHA, the elution peaks were observed at ~240 mM KCl, and for MllaSPARHA, at ~400 mM KCl. The corresponding peak fractions were collected, combined, and diluted with buffer containing 20 mM HEPES, pH 8.0, 5% glycerol, 2 mM EDTA to reduce the KCl concentration to 36 mM (28 mM for LheSPARHA). Subsequently, the samples were loaded onto a MonoQ 1 ml column (Cytiva) pre-equilibrated with buffer containing 20 mM HEPES, pH 8.0, 35 mM KCl (25 mM for LheSPARHA), 5% glycerol and 2 mM EDTA. After washing, the complexes were eluted with a KCl gradient from 35 mM (25 mM for LheSPARHA) to 800 mM in buffer containing 20 mM HEPES, pH 8.0, 5% glycerol and 2 mM EDTA. The elution profile revealed two peaks for AcaSPARHA (~200 and ~400 mM KCl) and LheSPARHA (~80 and ~450 mM KCl). For MllaSPARHA, a single peak at ~220 mM KCl was observed. Analysis of fractions by SDS-PAGE and mass-spectrometry showed that the peaks at ~200 and ~220 mM KCl corresponded to complexes of AcaSPARHA and MllaSPARHA, respectively. In the case of LheSPARHA, the peak at ~80 mM KCl corresponded to LheSPARHA, and the peak at ~450 mM KCl contained only LheHNH-APAZ. The fractions containing AcaSPARHA, MllaSPARHA, LheSPARHA, and LheHNH-APAZ were concentrated with an Amicon 30 kDa. The proteins were stored in a buffer with 50% glycerol and 2 mM DTT for further experiments.
Analysis of in vitro activities of SPARHA
For the analysis of nuclease activity of SPARHA on linear collateral substrates, 500 nM SPARHA (MllaSPARHA, AcaSPARHA, or their CD mutants, as well as LheSPARHA or LheHNH-APAZ) were incubated with 200 nM gRNA (unless otherwise indicated) in a buffer containing 20 mM HEPES, pH 8.0, 5 μg/ml BSA, 0.4 mM DTT, 5 mM Mn2+ (unless otherwise indicated), 5 mM KCl for AcaSPARHA and 100 mM KCl for MllaSPARHA or LheSPARHA for 15 min at 30 °C (unless otherwise indicated). Then, 200 nM complementary tDNA (unless otherwise indicated) was added, and after incubation for 10 min at 30 °C, 100 nM collateral substrate (ssDNA, RNA, dsDNA with pre-annealed strands) was added. After 1 h (or other time intervals as indicated in the Figures), the reaction was stopped by adding an equal volume of loading buffer containing 8 M urea, 20 mM EDTA, and 2x TBE. The reaction products were separated by 19% urea PAGE and visualized by SYBR Gold staining or fluorescence scanning (FAM for RNA substrates, HEX for dsDNA substrates) using a Typhoon FLA 9500 scanner (GE Healthcare). In the case of plasmid or genomic DNA as a substrate, the same protocol was used, except that the concentration of plasmid DNA was 2 nM and the amount of genomic DNA was 100 ng per 10 μl sample, and the reaction was stopped by adding 1/10 volume of loading buffer containing 50% glycerol, bromophenol blue, SYBR Gold (0.2 μl per 200 μl), and 0.1% SDS. The reaction products were separated on a 1% agarose gel and visualized by SYBR Gold staining.
To analyze the multi-turnover cleavage and substrate preferences of the SPARHA complex, 500 nM MllaSPARHA was incubated at 30 °C with 500 nM gDNA and tDNA as described above. Subsequently, 5 μM of various collateral substrates (DNA, RNA, or dsDNA of identical sequences) were added. The reaction was stopped as described above at specific time points (indicated in Supplementary Fig. 2b). Reaction products were separated by 19% urea PAGE and visualized by fluorescence scanning (FAM for the RNA substrate) or phosphorimaging for ssDNA and dsDNA (labeled with 32P at the 5’-end) using a Typhoon FLA 9500 scanner (GE Healthcare). See Supplementary Table 2 for oligonucleotide sequences.
Fluorescence assay
The AcaSPARHA complex (1 μM) was incubated with ori-gRNA-20nt (1.3 μM) for 15 min at 30 °C, then cooled on ice for 5 min. The pre-annealed collateral substrate dsDNA-32nt (1 μM) labeled with the ROX fluorophore and BHQ2 quencher was added on ice. Subsequently, ori-tDNA-20nt was added at various concentrations (as indicated in the legend to Supplementary Fig. 4b) on ice. Aliquots (15 μl) were transferred into a pre-chilled 384-well plate (CORNING, low volume), which was placed into a CLARIOstar reader. The incubation was carried out at 30 °C, and fluorescence measurements were taken every 5 min using Enhanced Dynamic Range.
Analysis of the effects of SPARHA on plasmids and phages
To analyze the effects of SPARHA expression on cell growth depending on the presence of interfering plasmids, E. coli BL21(DE3) cells were transformed with empty pBAD or pBAD_SPARHA plasmids (AcaSPARHA; LheSPARHA; or MllaSPARHA, including its mutants; all AmpR) and a second plasmid, either pCDFDuet-1 with the GST gene insertion (SmR) or pACYC184 (CmR). For analysis of growth curves, overnight cultures (grown with the corresponding antibiotics and 0.5% glucose) were diluted 75-fold in LB containing Amp, and 200 μl of the diluted culture was mixed with 40 μl of L-Ara to a final concentration of 0.2%. Subsequently, the cells were incubated at 25 °C (or 31.5 °C) with shaking in a 48-well plate using a CLARIOstar reader, with OD600 measured every 10 min. To determine the number of viable cells, samples were collected from the plate (after 8 h of incubation at 25 °C or after 10.5 h of incubation at 31.5 °C), and 10-fold serial dilutions were prepared and plated on LB agar containing Amp or the antibiotic corresponding to the interfering plasmid, with the addition of 1% glucose to suppress SPARHA expression. The number of colonies was counted after overnight incubation of the plates at 37 °C. For the experiment shown in Supplementary Fig. 5f, the protocol described above was used, except that antibiotics were not added to the overnight cultures or during the dilution of overnight cultures and after collecting samples from the 48-well plate, their 10-fold serial dilutions were plated on LB agar plates either without antibiotics or with the antibiotics corresponding to the plasmids.
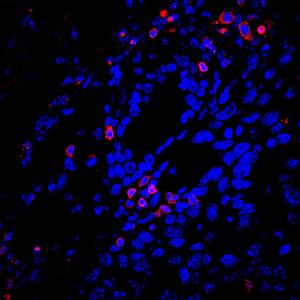
For cell staining, E. coli BL21 (DE3) containing empty pBAD, MllaSPARHA, and CD mutant of MllaSPARHA in the presence of the second plasmid pCDF were collected from a 48-well plate after 7.5 h of incubation at 25 °C with shaking in a CLARIOstar reader. The cells were pelleted by centrifugation, washed with 0.9% NaCl, and then resuspended in 0.9% NaCl to OD600~2.5. An equal volume of a mixture of SYTO9 (excitation/emission maxima: 485/500 nm) and SynaptoProbe Red (analog of FM4-64; excitation/emission maxima 515/640 nm) in 0.9% NaCl was added to the cells to final concentrations of 1 μM each. After 10 min of incubation at room temperature, 4 μl of the stained cells were visualized using a ZEISS LSM 900 confocal laser scanning microscope (Carl Zeiss) with excitation wavelength of 488 nm for SYTO9 and for SynaptoProbe Red, and images were taken using a 63× oil immersion objective. The experiment was performed in three biological replicates, with 5–10 fields per replicate.
To identify the effects of SPARHA on phage infection, the EOP was determined. E. coli BL21(DE3) cells were transformed with either empty pBAD or MllaSPARHA (WT or CD). Overnight cultures were diluted 30-fold (200 μl into 6 ml) in LB containing Amp and 0.2% L-Ara, and then incubated at 30 °C with shaking until OD600 ~ 0.5. Then 0.5 ml of cell suspension was mixed with 32 ml of 0.4% top agar containing 10 mM MgCl2, 5 mM CaCl2, Amp and 0.1% or 0.2% L-Ara (for experiments at 25 °C and 30 °C, respectively), and put onto plates containing 1.5% bottom agar (without any additives). PFU determination was performed by spotting 10 μl of 10-fold serial dilutions of phages HK140, HK243, mEp243, HK578, HK75, P1, λ, T7, T5, T6, HK225, HK106 and HK446. The plates were incubated at 25 °C or 30 °C and photographed after 14 h and 18 or 20 h of incubation. All incubation times are indicated above the plates in the figures. The PFU numbers were counted after 18 h for phages HK140, HK243, and mEp243, and after 20 h for phage HK578. In the case of overgrown phage spots, observed with SPARHA-sensitive phages, the PFU numbers were counted using the last phage dilution in which the spot remained visible. The number of phage particles in this spot was conventionally taken to be 100.
Analysis of small RNAs associated with SPARHA
To isolate small RNAs associated with MllaSPARHA in the presence of the interfering plasmid pCDF, overnight culture of E. coli BL21(DE3) cells transformed with pBAD_MllaSPARHA and pCDF-duet plasmids was added to LB medium containing Amp (150 μg/ml) and incubated at 30 °C with shaking for 1.5 h. Subsequently, L-Ara was added to a final concentration of 0.2%, and the culture was grown at 30 °C with shaking for 6.5 h. The cells were harvested by centrifugation (20 min, 5000 rpm, 4 °C), resuspended in lysis buffer (20 mM HEPES, pH 8.0, 1 M KCl, 10 mM K2HPO4, 2 mM PMSF, 2 mM DTT) and lysed by sonication. The supernatant was loaded onto a HisTrap HP 1 ml column (Cytiva) pre-equilibrated with buffer (20 mM HEPES, pH 8.0, 1 M KCl, 5% glycerol). After washing, the protein complex was eluted with the same buffer containing 300 mM imidazole. The eluate was concentrated using an Amicon 30 kDa device and loaded into 15% urea PAGE. The gel was visualized using SYBR Gold staining with a UV transilluminator, and the small RNAs were extracted from the gel, followed by overnight elution in 0.4 M NaCl.
For isolation of small RNAs associated with AcaSPARHA in the presence of the interfering plasmid pCDF, overnight culture of cells transformed with pBAD_AcaSPARHA and pCDF-duet plasmids was added to LB medium containing Amp (150 μg/ml) and 0.1% L-Ara, then grown for 8 h at 30 °C with shaking. The cells were harvested by centrifugation, resuspended in lysis buffer (20 mM HEPES, pH 8.0, 0.5 M KCl, 10 mM K2HPO4, 2 mM PMSF, 2 mM DTT) and lysed by sonication. Protein isolation of AcaSPARHA was performed as described above, including heparin and MonoQ chromatography steps, with the exception that EDTA was not added to the eluate after Ni-chelating chromatography and was excluded from all buffers. The peak of AcaSPARHA associated with small RNAs was eluted at ~370 mM KCl from the MonoQ column. The collected fractions were used for RNA isolation via phenol-chloroform extraction.
In both cases, small RNAs were precipitated using 3 M ammonium acetate, 96% ethanol, and glycogen, ethanol-washed, dissolved in water, phosphorylated using ATP and T4 polynucleotide kinase (NEB), and again precipitated, resulting in the final purified small RNAs. RNA libraries were prepared using the NEBNext Small RNA Library Prep Set for Illumina (Multiplex Compatible). Both RNA libraries (two biological replicates for MllaSPARHA and AcaSPARHA) were sequenced using Illumina HiSeq2500 (single-end 50-nucleotide reads). The list of sequenced small RNA libraries with corresponding accession numbers is shown in Supplementary Table 3.
Raw reads were processed to remove adapters using Trim Galore v.0.6.10, and the sequences were trimmed to a length of 15 to 25 nucleotides with Seqtk v.1.4-r122. Quality control before and after trimming was performed using FastQC v.0.12.1. The resulting libraries were mapped to the E. coli BL21(DE3) genome (NC_012971.2) and plasmids (pBAD_MllaSPARHA or pBAD_AcaSPARHA and pCDF) using STAR v.2.7.11b (with 97% alignment identity and intron exclusion, MatchNmin 14). Primary reads were extracted and used to generate sequence logos (16–25 nt or ≥20 nt in length) with WebLogo. Read counting of primary reads mapped to genes was performed using featureCounts v.2.0.8 (-s 1 for sense strand, -s 2 for antisense strand). The resulting values were normalized using RPKM, and distribution plots were created using custom Python scripts.
Mapping of the sites of DNA cleavage by SPARHA
To analyze the cleavage specificity, MllaSPARHA (500 nM) was preincubated with UA-gRNA-18nt (200 nM) in a reaction buffer (20 mM HEPES pH 8.0, 100 mM KCl, 5 μg/ml BSA, 2 mM DTT, 5 mM MnCl2) for 15 min at 30 °C. Control sample without MllaSPARHA was obtained in parallel. Complementary target DNA (200 nM) was added to the reaction mix, and the samples were incubated for 10 min at 30 °C. Genomic DNA (80 ng) was added (purified from E. coli BL21(DE3) transformed with pBAD_Mlla and pCDF-Duet), and the cleavage reactions were performed for 10 min at 30 °C. The samples were mixed with 10x loading buffer (50% glycerol, 0.1% SDS, SYBR Gold), chilled on ice, and aliquots were analyzed by 1 % agarose gel electrophoresis to confirm DNA smearing in the reaction containing MllaSPARHA. DNA from the remaining samples was treated with phenol-chloroform and ethanol-precipitated. For library preparation, DNA ends were blunted using the DNA end repair mixture and adapter sequences described previously68. The samples were treated with 2 μl of 3 U/μl T4 DNA polymerase (NEB), 2.5 μl of 10 U/μl T4 polynucleotide kinase (NEB), 0.5 μl of 5 U/μl, Large Klenow Fragment of DNA Polymerase I (NEB) in total volume of 50 μl containing 5 μl 10x T4 DNA ligase buffer (NEB) and 2.5 μl 10 mM dNTP mix (Thermo Fisher Scientific, R0191) for 45 min at 25 °C. DNA was purified with Agencourt AMPure XP beads and eluted with 25 μl of 10 mM Tris-HCl (pH 8.0). To perform the A-tailing reaction, the DNA samples were supplemented with 15 μl of MiliQ-grade water, 5 μl of 10x NEBuffer 2.1 (B6002S), 2.5 μl of 10 mM dATP and 2.5 μl of 5 U/μl Klenow (exo-) (NEB). The reactions were carried out for 45 min at 37 °C, and the enzyme was heat-inactivated by incubation for 20 min at 65 °C. DNA was purified using Agencourt AMPure XP beads and eluted with 40 μl of 10 mM Tris-HCl (pH 8.0).
Detector adapter with a 3’-dT overhang was generated by annealing Detector_adapter_dir and phosphorylated Detector_adapter_rev oligonucleotides as described previously68. The resulting adapter (2.5 μl of 10 μM solution) was ligated to the purified 3’-dA overhanged cleavage products in 50 μl samples containing 5 μl of 1 U/μl T4 DNA ligase and 5 μl of 10x T4 DNA ligase buffer (NEB) for 16 h at 22 °C. The ligated DNA was purified using Agencourt AMPure XP beads and eluted with 28 μl of 10 mM Tris-HCl (pH 8.0). Next, sequencing primers were added using the NEBNext Ultra II FS DNA Library Prep Kit for Illumina. The resulting samples were amplified with dual index primers (NEBNext Multiplex Oligos for Illumina) and purified with Agencourt AMPure XP beads. PCR products were sequenced on a NovaSeq 6000 System (Illumina) in the 100 bp paired-end mode. The list of sequenced DSB libraries with corresponding accession numbers is shown in Supplementary Table 3.
Raw reads were processed to filter by quality using Trim Galore v.0.6.10; quality control before and after filtering was performed with FastQC v.0.12.1. Subsequently, reads containing the detector adapter sequence at the 5’-end were extracted from libraries for further analysis (for MllaSPARHA and control), and the adapter sequence was further removed using custom Python scripts. The resulting library was mapped to the genome (NC_012971.2) and plasmids (pBAD_MllaSPARHA and pCDF-Duet) using STAR v.2.7.11b (with 97% alignment identity and intron exclusion).
To construct sequence logos, the first 5 nucleotides of primary reads (distance from +1 to +5) and 6 nucleotides to the left of the read start (distance from −5 to 0) (Fig. 5j and Supplementary Fig. 14b) were extracted from the reference genomic and plasmid sequences using a custom Python script. Sequence logos of the resulting FASTA files were generated using WebLogo. To analyze the distribution statistics of distances between the 5’-ends of reads on the plus and minus strands in the genome, a custom Python script based on a binary search algorithm was used. The resulting statistics were visualized with a custom Python script. To construct a sequence logo for pairs with a distance of 1, reads from the identified pairs were extracted into a separate BAM file, and the logo was generated as described above.
Size-exclusion chromatography and SEC-MALS
The absolute molar masses and the oligomeric state of the apoforms of AcaSPARHA and MllaSPARHA were determined by size-exclusion chromatography (SEC) coupled to multi-angle light scattering (MALS). Proteins were pre-incubated for at least 15 min at 10 °C in the Vialsampler (G7129A, Agilent) and loaded on a Superdex 200 Increase 10/300 column (Cytiva) operated using an Agilent 1260 Infinity II chromatography system equipped with a 1260 Infinity II WR diode-array detector (G7115A, Agilent), a miniDAWN detector (Wyatt Technology) and a refractometric detector (G7162A, Agilent, operated at 35 °C). The column buffer contained 20 mM HEPES-KOH pH 8.0 with 100 mM KCl, and the flow rate was 1 ml/min. Post-column filters were not used. The elution profiles were followed by absorbance at 280 nm and changes of the refractive index (RI), as well as static light scattering at three angles. The data were collected and processed in Astra 8.0 software (Wyatt Technology) using a refractometric detector as a concentration source (dn/dc was taken equal to 0.185 for each protein) and 250,000 attenuation of the output RI signal. Normalization of the static light scattering signals at different angles detected by miniDAWN was done using a pre-run of a BSA standard (Wyatt). The data were analyzed with Astra 8.0 (Wyatt Technology, USA) using dn/dc = 0.185. The polydispersity index (the Mw/Mn ratio) of the protein peaks was around 1.000.
To screen for the effects of the metal ion, gRNA and tDNA on the apparent size and oligomeric behavior of protein samples, the samples were pre-incubated at 30 °C within the Vialsampler and injected in 28 μl aliquots onto a Superdex 200 Increase 5/150 column (Cytiva) operated at a 0.45 ml/min flow rate. The column was pre-equilibrated with 20 mM HEPES-KOH pH 7.0, containing 50 mM KCl and 0.1 mM MnCl2. In the presence of gRNA (ori-gRNA-20nt) and tDNA (ori-tDNA-20nt, Supplementary Table 2), both WT and CD AcaSPARHA displayed a large peak in the void column volume, which for Superdex 200 Increase corresponded to ≥600 kDa. To assess more specifically what large size this species corresponds to, we performed SEC-MALS using a Superose 6 Increase 10/300 column (Cytiva), which has an exclusion limit of ≥5 MDa. Column buffer was 20 mM HEPES-KOH pH 7.0 containing 50 mM KCl and 0.1 mM MnCl2. The flow rate was 1 ml/min. The data were analyzed in A 8.0 (Wyatt Technology, USA) using dn/dc = 0.185 and the extinction coefficient ε (0.1%) at 280 nm of 1.29 (mg/ml)–1cm–1. The protein conjugate analysis implemented in Astra 8.0 (Wyatt) was used to assess mass contributions from protein and bound nucleic acid components in the elution peaks, using the dn/dc and ε (0.1%) values indicated above for Aca, dn/dc = 0.17, ε (0.1%) at 280 nm of 15.8 (mg/ml)−1cm−1 for the gRNA (ori-gRNA-20nt), or dn/dc = 0.17, ε (0.1%) at 280 nm of 10 (mg/ml)–1 cm–1 for the RNA/DNA duplex (ori-gRNA-20nt/ori-tDNA-20nt).
Statistical analysis
Statistical analysis of the experimental data was performed with the statannotations Python package. P-values were calculated by the t-test for independent samples and corrected using either the Bonferroni or Holm-Bonferroni correction, as indicated in the figure legends.
Transmission electron microscopy with negative staining
To visualize SPARHA complexes with gRNA and tDNA by negative-stain TEM, 2.5 μM MllaSPARHA, AcaSPARHA, or their mutants were incubated with 2.5 μM gRNA in a buffer containing 20 mM HEPES (pH 7.0), 100 μM Mn2+, and either 100 mM KCl for MllaSPARHA or 50 mM KCl for AcaSPARHA for 10 min at 30 °C. Subsequently, 2.5 μM complementary tDNA was added, and incubation was continued for an additional 2.5 min. Substrate plasmid DNA (300 ng per 20 μl sample) was added simultaneously with tDNA, when indicated. Control samples of apo-SPARHA without gRNA and tDNA were prepared in parallel. To prepare the samples for TEM, 3 µl of sample suspension was applied to a carbon substrate film (EMCN, China) hydrophilized with glow discharge. Negative staining was performed using 1% uranyl acetate for 30 s.
TEM imaging was carried out using JEM-1400 120 kV transmission electron microscope (JEOL, Japan) equipped with Rio 9 camera (Gatan, USA). For 3D reconstruction, image series were acquired with JEM-2100 200 kV transmission electron microscope (JEOL, Japan) equipped with a DE-20 detector (Direct Electron, USA). Automated data collection was performed with SerialEM69 at a defocus of –1.8 µm. For each sample, 600–1200 images were acquired. The contrast transfer function (CTF) was estimated with CTFFIND470. 3D reconstruction was performed in cryoSPARC71. Individual helical filament segments (particles) were picked using template-based selection based on manually selected particles.
Structural heterogeneity, particularly the presence of different helix types within the MllaSPARHA protein complex, was resolved through multiple rounds of 2D classification. Helix parameters were initially determined via asymmetric helical refinement and subsequently refined in the final round of helical refinement. The resulting reconstructions were generated from 3000–12,000 particles, depending on the sample, and achieved a resolution of 16 Å based on the Fourier Shell Correlation (FSC) 0.143 criterion. This resolution is consistent with the limitations of the negative staining method.
Structural modeling with AlphaFold
All models were predicted by AF3. The model of the MllaSPARHA HNH tetramer with dsDNA was manually adjusted using PyMol, based on the interactions of residues between HNH dimers in the structure of the Cap5 tetramer (PDB ID:8FMF). The coordinates of the AF3 models generated during this study are provided as a zip archive (Supplementary Data 1 with SPARHA AlphaFold3 models). The pLDDT scores and PAEs for each model are provided in Supplementary Data 2. For building the models of filaments of AcaSPARHA and MllaSPARHA containing gRNA and tDNA, the tetramer models were copied and aligned head-to-tail by the MllaSPARHA or AcaSPARHA chains using UCSF ChimeraX.
Purification of AcaSPARHA protein for cryo-EM and grid preparation
Wild-type AcaSPARHA complex was expressed in E. coli BL21(DE3) as described above. Cells collected from 2 L of the LB culture were lysed by sonication in buffer containing 20 mM Tris-HCl (pH 8.0), 1 M NaCl, 5% glycerol, 2 mM DTT, and 2 mM PMSF. The lysate was clarified by centrifugation and loaded onto a Ni-Sepharose column pre-equilibrated with 20 mM Tris-HCl (pH 8.0), 0.5 M NaCl and 5% glycerol. After washing with equilibration buffer and 20 mM imidazole, the protein was eluted with 300 mM imidazole. EDTA (5 mM final) was added to the elution fractions, incubated for 20 min, and the fractions were diluted sequentially 4 times with equilibration buffer and 20 times with low-salt buffer (20 mM Tris, pH 8.0, 5% glycerol, 2 mM EDTA). The diluted fractions were loaded onto a heparin-Sepharose column pre-equilibrated with 20 mM Tris-HCl (pH 8.0), 30 mM NaCl, 5% glycerol, and 2 mM EDTA. Elution was performed using a linear salt gradient (30–600 mM), with AcaSPARHA eluting at ~240 mM NaCl. Pooled fractions were diluted to 36 mM NaCl and applied to a Hi-Trap Q anion exchange column equilibrated with 20 mM Tris-HCl (pH 8.0), 30 mM NaCl, 5% glycerol, and 2 mM EDTA. A linear NaCl gradient (35–600 mM) resolved two peaks with AcaSPARHA eluted at ~200 mM NaCl and impurity (likely corresponding to an HNH-APAZ monomer) eluted at ~400 mM NaCl. Fractions containing AcaSPARHA were concentrated using a 10 kDa Amicon filter. To prepare the complex with gRNA and tDNA for cryo-EM specimen preparation, AcaSPARHA (20 μM) was mixed with 20 μM gRNA in 20 mM Tris-HCl (pH 7.0), 50 mM NaCl, and MnCl2 (100 μM) and incubated for 10 min at 30 °C. Target DNA (25 μM) was added and incubated for ≤2.5 min at 30 °C. Samples (3.5 µL) were applied to glow-discharged UltraAuFoil 1.2/1.3 grids, blotted, and plunge-frozen in liquid ethane using a Vitrobot Mark IV (FEI) at 4 °C (100% humidity).
Cryo-EM data acquisition and processing
Cryo-EM data were collected on a 200 keV Talos Arctica TEM (Thermo Fisher) with a Falcon 4 detector (Gatan) at the Pennsylvania State University Huck Institutes Cryo-Electron Microscopy Core Facility (Supplementary Fig. 10a). The defocus range was –1.0 to –2.0 µm, and the magnification was 150,000× in electron counting mode (pixel size = 0.944 Å/pixel). Fifty frames per movie were collected with a nominal dose of 50 e-/Å2 (Supplementary Table 4).
The data were processed using cryoSPARC (Supplementary Fig. 10b)71. The movies were aligned and dose-weighted using Patch-motion correction and Patch-CTF estimation, followed by discarding low-quality micrographs that had large motions or poor CTF resolution through a manually curated exposure job. 159,273 particles were picked by filament tracer (Filament diameter = 160 Å, separation distance = 0.5, minimum filament length = 480 Å, Supplementary Fig.10a) followed by extraction (box size = 600 pixel), and subjected to 2D classification (Supplementary Fig. 10c) to remove poorly populated classes. The Initial 3D volume was made by ab initio reconstruction, and heterogeneous refinement was used to remove bad particles. A class representing AcaSPARHA filament was selected and refined to generate a homogenous density map at 3.40 Å resolution. Mask was created around the 3D volume corresponding to the AcaSPARHA tetramer for local refinement to improve density map at 3.38 Å resolution. The nominal resolution of the cryo-EM map was estimated by 0.143 gold standard FSC cut off (Supplementary Fig. 10d). The 3D refinement with helical symmetry (helical refinement in cryoSPARC) showed helical symmetry with a twist angle of 92.44° and a rise size of 103.25 Å (tetramer as an asymmetric unit), but its resolution (4.48 Å) was lower than the resolution of reconstruction without helical symmetry, due to its curvature and heterogeneity.
Model building and structure refinement
The initial model was built by AlphaFold72 as described above. Modeling errors found in the AcaSPARHA were corrected by using Coot73. The structure was real-space refined by using rigid-body refinement, secondary structure, Ramachandran, and rotamer in Phenix74 (Supplementary Table 4).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.