NeoRHEA clinical study
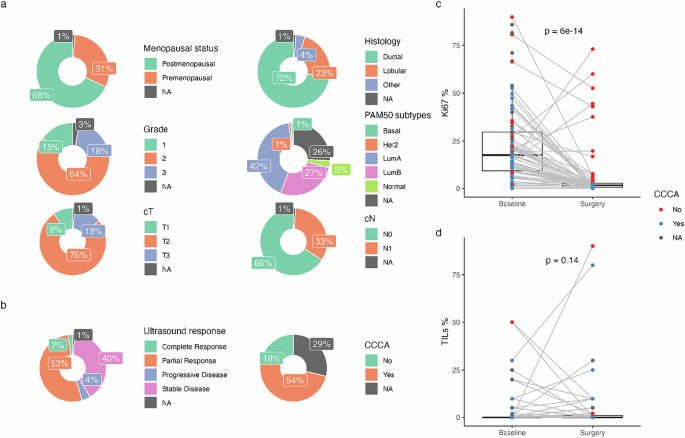
NeoRHEA (clinical trials gov identifier NCT03065621) is an open-label, single-arm, phase 2 study, including patients with ER + /HER2- early breast cancer candidates for neoadjuvant therapy. The study protocol is provided as supplementary material. All patients have been treated with 4 cycles of endocrine treatment and palbociclib, plus three to seven days of the same treatment (bridge period), so that they were on treatment at the time of surgery. A total of 100 patients were recruited between July 2017 and March 2019 in 5 hospitals in Belgium. Out of 100 patients, two have withdrawn consent, and one had bone metastases at the time of diagnosis and thus were excluded from the analyses. All remaining 97 patients have provided informed consent to participate in the study and related translational research. The study was approved by the Institut Bordet Ethical Committee. The full study protocol is provided as supplementary material. A CONSORT flow diagram for the NeoRHEA study, including the bulk RNA, and single-nuclei RNA and ATAC-seq analyses, can be found in Supplementary Fig. 11.
Criteria for ultrasound response
The main method for evaluation of ultrasound (US) response was bi-dimensional (longest diameter x greatest perpendicular diameter of primary tumor) ultrasound, performed before and after study treatment. Response criteria were those of the World Health Organization as evaluated through US32. Patients were evaluated at the end of the 4-month neoadjuvant treatment and were classified as having tumors with complete response (CR), partial response (PR), stable disease (SD), or progressive disease (PD). Patients with CR or PR were classified as responders, whereas patients with SD or PD as non-responders.
Ki67 and sTILs central pathology review
Tumor biopsies have been collected at baseline (before treatment) and after 4 months of treatment (at surgery). Histopathological assessment of all formalin-fixed parrafin embedded (FFPE) tumor samples collected before and after neoadjuvant treatment were centrally performed, including: % tumor cellularity, Ki67 status by immunohistochemistry (IHC) and stromal Tumor Infiltrating Lymphocytes (sTILs). Ki67 and PanCytokeratine (PanCK) staining were performed using the monoclonal mouse anti- Human Ki67 antigen Clone MIB-1 (Dako Omnis) and monoclonal mouse anti-human cytokeratin clone AE1/AE3 Ready-To-Use (Dako Autostainer/Autostainer Plus), respectively. Briefly, PanCytokeratine staining was used to select only the Ki67 staining within the tumor cells, excluding Ki67-expressing lympho- cytes. In order to do so, adjacent sections stained with each antibody were scanned using a digital slide scanner Nanozoomer (Hamamatsu) that converts glass slides into high-resolution digital data by high-speed scanning, and subsequently aligned using Visiopharm (Visopharm®), an histopathology image analysis software for can- cer diagnostics and research with Artificial Intelligence (AI) and deep learning technologies in Augmented Pathology®.
Of note, this procedure allows us to have a precise percentage (%) of Ki67 expression on a scale of 1–100%. Therefore, it allowed us to evaluate the presence or absence of Complete Cell Cycle Arrest (CCCA) in our patients’ tumors, defined as centrally reviewed Ki67 ≤ 2.7% at surgery. The percentage of sTILs was centrally evaluated using Hematoxylin and Eosin (H&E) slides according to the guidelines of the TILs working group33.
Bulk RNA sequencing
RNA extracted from frozen tumor and FFPE samples with at least 25% centrally- assessed tumor cellularity was processed for RNA-sequencing. Strand-specific cDNA libraries were constructed using the NEBNext Ultra directional RNA library Preparation Kit for Illumina paired-end sequencing on a NovaSeq 6000 instrument (Illumina). Briefly, starting from 200 ng of total RNA, rRNA was depleted using the Ribo-Zero Magnetic Kit (Epicenter, Wisconsin) according to the manufacturer’s instructions. Library quality control and quantification was performed using the Fragment Analyzer (Advanced Analytical) and the QUBIT® (Invitrogen). Clusters were generated in a cBot Cluster Generation System (Illumina) using the Kits TruSeq PE Cluster Kit v3-cBot-HS and HiSeq PE Cluster Kit v4 cBot and were sequenced on the Illumina NovaSeq 6000 in 2 x 100bp paired-end mode. In total matched (baseline + surgery) FFPE from 18 patients, matched frozen from 31 patients and unmatched frozen baseline from 29 patients were sequenced. Salmon quantifying the expression of transcripts was used in mapping-based mode using a hg38 reference genome.
Single-nuclei sequencingMultiome Wet Lab
We used the 10x Chromium Single Cell Multiome ATAC + Gene Expression kit. Frozen optimal cutting temperature (OCT) embedded baseline (pre-treatment) biopsy and surgery tumor samples with at least 25% tumor cellularity were selected for single-nuclei (sn)RNA-seq and ATAC-seq analysis. Briefly, samples were processed into the single-nuclei dissociation with protocol C “Multiome” – adapted version of “Nuclei Isolation from Complex Tissues for Single Cell Multiome ATAC + Gene Expression Sequencing”. Nuclei were first isolated by multiple steps of centrifuging and lysis, then they were sorted, and lastly, they were permeabilized again by means of centrifuging and lysis. Then, extracted nuclei underwent snRNA-seq & ATAC-seq using Chromium Next GEM Single Cell. In order to retain optimal nuclei membrane quality, we modified the aforementioned protocol by using a 10X lysis buffer instead of a 1X lysis buffer for the isolation step. In total, samples from 37 patients were successfully processed (matched samples from 26 patients, unmatched baseline samples from 4 patients and unmatched surgery samples from 7 patients).
Sequencing
The sequencing of the samples was done at the BRIGHTcore Genomics core facility of Uni- versite Libre de Bruxelles (ULB). 10x Genomics-generated cDNA libraries were sequenced on the Illumina NovaSeq 6000 instrument. The sequencing metrics for the GEX was: Read 1-28 cycles, I7-10 cycles, I5-10 cycles, Read 2-90 cycles. The sequencing metrics for the ATAC was: Read 1-50 cycles, I7-8 cycles, I5-24 cycles, Read 2-49 cycles.
Bioinformatics methods
To arrive at count matrices for both RNA-seq and ATAC-seq data, various components of Cell Ranger ARC v2.0, provided by 10X Genomics, were used. More specifically, to demultiplex raw base call files generated by Illumina sequencers “cellranger-arc mkfastq” was used for both ATAC and RNA flow cells. Doing that resulted in a handful of fastq files for each sample, which were then used as an input to “cellranger-arc count” using the hg38 reference for both the RNA and ATAC assays. The latter trimmed and aligned all fastq files to the reference genome in use and then performed filtering, peak calling and counting for both the ATAC and RNA molecules. Doing these steps, we produced count matrices for both the RNA-seq and ATAC-seq samples. Having arrived at count matrices for each cell, RNA counts were processed with the standard Seurat (version 4.3) workflow (normalize data, find variable genes, scale data, PCA, find cell clusters using sNN with resolution set to 0.5 and lastly run dimensionality reduction projections such as t-SNE and UMAP) while ATAC counts were processed again with Signac by finding the most frequently observed peaks, calculating singular value decompositions and dimensionality reduction projections. Before normalizing any of the produced count matrices, we first applied some quality control filters to discard bad-quality nuclei. Having predicted cell type annotation for the RNA partition of our data, the snATAC-seq had predicted annotations transferred from the snRNA-seq dataset. Cell types were identified by expression patterns of known in the literature biomarkers and tumor cells were identified by profiling the copy number aberrations of epithelial cells via infercnv. Concerning the snATACseq, peaks were called for all cells after quality control filtering via MACS2. After quantifying reads in the snATAC-seq data, we were able to assess the activity of transcription factor motifs by cell, utilizing chromVAR using motifs from the JASPAR 2022 transcription factor database. After this, differential tests comparing genes and peaks between baseline and surgery conditions were performed. These included differential gene expression (DGE) analyses, differential peak accessibility analyses, differential motif activation analyses, as well as subsequent over gene set enrichment analyses (GSEA). Lastly, we performed an analysis linking peaks with genes that were concurrently expressed and accessible.
Quality control for snRNA-seq & ATACseq
Samples with less than 400 or more than 16,000 nuclei were immediately excluded, as this indicates suboptimal wet lab processing of the respective samples. Samples passing that filter were included in the rest of the droplet quality control process. Droplets not identified as nuclei by cellranger were excluded. For the snRNAseq, regarding the number of unique molecular identifiers (UMIs), droplets with less than 400 or more than 25,000 were excluded and regarding the number of genes, droplets with less than 200 or more than 10,000 were also excluded. We computed mitochondrial and ribosomal genes percent for each of the droplets, and those exceeding 2% of mitochondrial genes or 1% of ribosomal genes were excluded. For the snATAC-seq droplets with less than 1000 or more than 100,000 numbers of UMIs were excluded. Peaks were then called inputting fragments of nuclei from all samples together via MACS2 so that the peak set is common for all samples, and peaks with width less than 20 base pairs or more than 10,000 base pairs were excluded.
Cell type annotation via markers and copy numbersHigh level labels
Initially, we spit our cell population into three categories – epithelial, stromal, and immune – based on respective markers. For epithelial cells we used Keratin 19 (KRT19), Estrogen Receptor 1 (ESR1) and Ankyrin Repeat Domain 30 A (ANKRD30A); for stromal cells we used Platelet And Endothelial Cell Adhesion Molecule 1 (PECAM1), Fibroblast Activation Protein Alpha (FAP) and Actin Alpha 2, Smooth Muscle (ACTA2) and lastly for immune cells we used CD68 Molecule (CD68), Membrane Spanning 4-Domains A1 (MS4A1) and Protein Tyrosine Phosphatase Receptor Type C (PTPRC). Cells were labeled based on their cluster associations with each of the markers, meaning that for each cluster, we carefully inspected which of the selected markers were mostly expressed and then manually labeled all cells of the respective cluster to the cell type whose markers were most expressed.
Tumor vs normal cells
Having identified the three major cell type populations, we then profiled copy number aberrations of the epithelial cells specifically to distinguish the malignant cells from the normal epithelial ones. To do that we created a reference set of 500 immune and stromal cells that were thought to be copy number neutral. For each patient, we selected proportionally in relation to the rest of the patients the least variable in terms of expression of immune and stromal cells, resulting in 500 cells in total. Then, for each sample, we separated the epithelial cells from the rest and profiled their copy number status using the previously generated set of reference cells via infercnv in subclusters analysis mode, without clustering by groups and by using an HMM (hidden Markov model) model and denoising technique. Having profiled the copy number status of the epithelial cells by employing a KNN (k nearest neighbor) classification strategy for each sample, we compared the copy number profile of each cell against the profile of the reference set of cells and the profile of the top 5% most aberrant cells of the same sample. Cells whose copy number profile resembled most the one of the reference set of cells were labeled as normal epithelial cells, while the rest were labeled as tumor cells.
Stromal and Immune cells subclassification
Stromal cells were further labeled as either fibroblasts, myofibroblasts or endothelial cells again by using specific markers and leveraging the cluster information. More specifically, FAP was used to label fibroblasts, ACTA2 to label myofibroblasts, and lastly, PECAM1 to label endothelial cells. Immune cells were classified into T, B and myeloid cells by using CD3 Epsilon Subunit Of T-Cell Receptor Complex (CD3E), MS4A1 and CD68 respectively.
Immune T cells subclassification
All immune T cells were labeled as either CD4 or CD8 based on their expression of CD4 Molecule (CD4) and CD8 Subunit Alpha (CD8A). CD8 cells were further labeled as tissue resident memory (TRM) cells based on their expression of Integrin Subunit Alpha E (ITGAE) or CD103, and CD4 cells further labeled as FOXP3-positive regulatory cells.
Differential gene expression and gene set enrichment analyses
For bulk RNA sequencing data, DESEq2 was used for DGE analysis with fold change shrinkage as implemented in the lfcShrink function, of note for patients with both OCT and FFPE samples available in any of the time points, the sample with paired counterpart in the other time point was in use. To take into account the effect of samples originating from tissues with different conservation methods (OCT vs FFPE) the conservation method was added in the generalized model comparing expression of genes. In the single-nuclei cohort, after robustly set cell type labels for each of the cells included in our cohort, we then performed DGE analyses between baseline and surgery cells for each of the cell type subpopulations. For these analyses, we used the FindMarkers Seurat command with default settings the log-fold threshold, as well as the minimum percentage of cells expressing each gene to zero. To infer biological conclusions, using the DGE analyses, we subsequently performed GSEA using the Hallmark gene sets. For all the GSEA analyses, the clusterProfiler Bioconductor with the fgsea method was in use.
Differential peak accessibility analyses
To gain insights into changes in chromatin accessibility in our set of cells post-treatment, we performed differential peak accessibility analyses again for all identified cell types. The Seurat FindMarkers command was in use however, this time the test utilized logistic regression with the total number of fragments in each of the peaks added as a latent variable to mitigate the effect of differential sequencing depth on the results. Log-fold threshold was set to zero and the minimum percentage of cells with each of the peaks being accessible was set to 5%.
Gene set enrichment analysis of peaks
To perform a conventional GSEA analysis using the differentiated peak, there is no need to split each set of peaks into upregulated and downregulated peaks; however, a fold change of genes and not peaks is needed. Having both a set of differentiated genes and peaks, we select the genes residing in the differentiated peaks and use the fold change as calculated in the DGE analysis of these specific genes as an input to the GSEA analysis using the Hallmark gene set.
Multiplex immunohistochemistry PD-L1, PD-L2
Multiplex immunohistochemistry (mIHC) was performed in the context of an ongoing collaboration between Institut Bordet and Rejuveron. mIHC to stain for CK, CD45, Ki67, Programmed death-ligand 1 and 2 (PD-L1 and PD-L2) and 4’,6-diamidino-2-phenylindole (DAPI) was performed on 36 baseline and 33 surgery FFPE tumor samples from 37 patients. The markers included in this panel aimed to validate our snRNA-seq findings on the decrease in the proliferation of tumor and immune cells. Moreover, we wanted to study changes in the expression of the PDL1 and PDL2 ligands of the immune-checkpoint receptor programmed cell death protein 1 (PD1), which is the most clinically relevant checkpoint in cancer immunotherapy (Pardoll et al.34). Immunofluorescence images were first unmixed using InForm, and then subsequent cell quantifications were done using QuPath (version 0.5.1). Cell segmentation was performed by employing the stardist algorithm, filtering out cells with a detection probability lower than 50%. For each of the markers, a boolean classifier was trained, classifying each cell as positive or negative. All classifiers were combined in one composite classifier, with which the final classification was performed. Cells were labeled as tumor if they were classified as panCK positive and CD45 negative, immune if they were classified as panCK negative and CD45 positive and stromal if they were classified as panCK negative, CD45 negative and Ki67 negative.
Multiplex immunohistochemistry CD103, CD39
Multiplex immunohistochemistry (mIHC) was performed on FFPE tumor tissues collected at baseline and post-surgery from seven patients, selected based on their high tumor infiltrating lymphocyte (TIL) scores. The mIHC panel included: CD4, CD8, CD39, CD103, CK and Ki67 plus DAPI for nuclear staining. The markers selected in this panel aimed to validate our snRNA-seq findings on the decrease in T-cell proliferation and TRMs. CD39 was included in the panel based on published data showing that co-expression of CD103 and CD39 identifies a subset of tumor-reactive T cells in solid tumors35. Of note, the mIHC panel optimization was done according to the procedure we have previously detailed36 before processing the samples from the 7 NeoRHEA patients. Image acquisition was achieved using a Vectra PolarisTM (Akoya Biosciences®), and subsequent analysis and quantification was performed using InForm, phenoptR and PhenoptrReports (Akoya Biosciences®). Images were preprocessed for spectral unmixing to identify marker signals, followed by tissue segmentation to differentiate tumor and stroma regions using machine learning algorithms trained on manual annotations. Nuclear segmentation was performed using the DAPI staining. Phenotyping categorized cellular subpopulations of tumor cells (pan-cytokeratin or CK + ), CD4 + T cells, CD8 + T cells, and total T cells (CD4 + plus CD8 + ). Tissue-resident memory T cells (TRM) were identified as CD8 + T cells co-expressing CD103. Proliferating cells (tumor and immune) were identified by Ki67 positivity. CD39 expression was assessed across all subpopulations, with CD4 + and CD8 + T cells the principal positive subpopulations; however, a large proportion of vessels also stained positive, increasing the background for this marker.
FELINE trial
FELINE trial (clinicaltrials.gov identifier NCT02712723) randomized in a 1:1:1 ratio 120 post-menopausal women with node-positive or ›2 cm ER + /HER2- breast cancer to a 6-month neoadjuvant treatment with either letrozole alone or letrozole and intermittent administration of ribociclib or letrozole and continuous administration of ribociclib. snRNA-seq using 10X genomics was performed on tumor biopsies collected at baseline (pre-treatment), after 14 days of treatment and at the end of treatment (surgery). We analyzed snRNA-seq data from 23 patients with letrozole alone and 39 patients with letrozole and ribocilib combining the intermittent and continuous ribociclib schedule that we have retrieved from Griffiths et al., 2025. We only used data from tumor biopsies at baseline and surgery that represented similar timepoints with the NeoRHEA study. We used the count matrices and cell type annotations provided in ref. 11. We have performed DGE analyses between baseline and surgery cells for each of the cell type subpopulations. To infer biological conclusions, using the DGE analyses, we subsequently performed GSEA using the Hallmark gene sets as we did in the NeoRHEA study.
Statistics and reproducibility
The study primary objective was to develop biomarkers to predict resistance to neoadjuvant treatment of endocrine therapy and palbociclib using RNA sequencing of the baseline tumor biopsy. The following few assumptions were made: The overall response rate would be around 75%-80%. The resistance rate would be therefore around 20%-25%. According to simulations carried out by Dobbin et al.37, a sample size of at least 20 subjects in each class would be required to develop a binary predictor using high-dimensional data. Therefore, to get 20 subjects in the resistant group with an expected 22.5% resistance rate, around 90 evaluable subjects would be needed. To take into account a rate of 10% of non-evaluable subjects, 100 eligible subjects would be required for inclusion in the study. For all the analyses between baseline and surgery samples using either bulk RNA-seq or snRNA-seq and ATAC-seq data, as well as for all mIHC analyses, no statistical method was used to predetermine sample size.
NeoRHEA data availability
The bulk and single-nucleus RNA-seq data generated and analyzed in this study have been deposited in the European Genome-phenome Archive (EGA) under the accession number ID – EGAD50000002038. Access to the data is controlled and subject to approval by the Data Access Committee, composed of Michail Ignatiadis, Christos Sotiriou, and David Venet. There are no restrictions on eligibility for data access, provided that the submitted proposal is approved by the committee. For data access or related queries, please contact David Venet and Michail Ignatiadis. Data access requests are typically processed within 3–6 months, depending on the nature of the request. The data will remain available for the duration of the approved proposal.
Raw images from mIHC experiments and count matrices from bulk RNA-seq, and snRNA-seq, and ATAC seq data have been uploaded in Zenodo DOI 10.5281/zenodo.16760008. Source data are provided as Source Data files.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.